| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,401,662 – 8,401,754 |
| Length | 92 |
| Max. P | 0.999862 |

| Location | 8,401,662 – 8,401,754 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 75.50 |
| Mean single sequence MFE | -19.98 |
| Consensus MFE | -11.49 |
| Energy contribution | -11.69 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.58 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982297 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
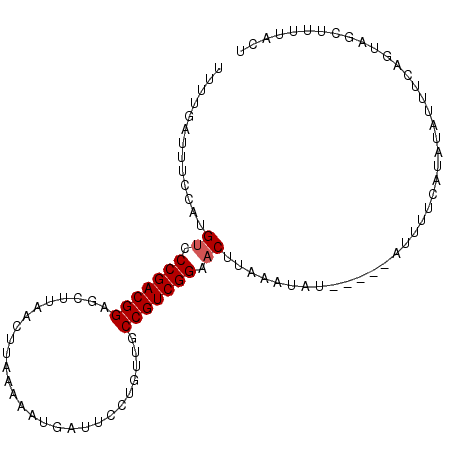
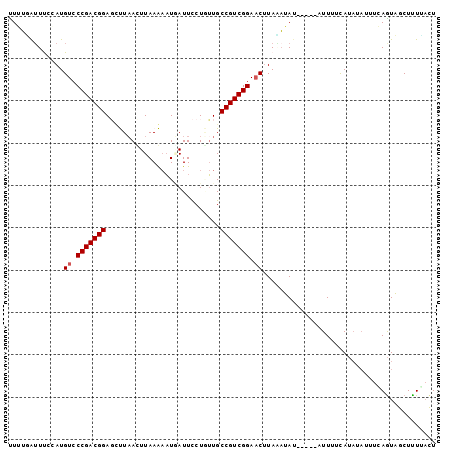
>2R_DroMel_CAF1 8401662 92 + 20766785 UUUGUGUUUUCAUGUCCCGACGGAGGUUUAUUUAAAAAUGAUUCCUAUUGCCGUCGGACCUUAGCUAU-----AUUUUCA----UUUCAAUAGCUUUUACU ...(((.......((((.((((((((.(((((....)))))..)))....)))))))))...((((((-----.......----.....))))))..))). ( -19.60) >DroSec_CAF1 12632 96 + 1 UUGUUAUUCCCAUGUCCCGACGGAGUUUAACUUAAAAAUGAUUCCUGUUGCCGUCGGAACUUAAAUAU-----AUUUUCAUAUAUUUCAGUAGCUUUGAUU ..((((((.....((.(((((((....((((...............))))))))))).))..((((((-----((....)))))))).))))))....... ( -18.26) >DroSim_CAF1 12577 96 + 1 UUGUAAUUCGCAUGUCCCGACGGAGUUUAAUUUAAAAAUAAUUCCUGUUGCCGUCGGAACUUAAAUAU-----AUUUUCAUAUAUUUCAGUAGCUUUUACU ..((((...((.....(((((((.....((((.......)))).......))))))).(((.((((((-----((....)))))))).))).))..)))). ( -21.20) >DroEre_CAF1 14809 101 + 1 CUUUGAGUUCGAUGUCCCGACGGAUCCUAACUUAAGAAUUAUUUCUUUUGCCGUCGGAACUUAUAAAAUUUUAAUUUUCAUAUAUUUGAGUUGCUCUCUCU ....(((..((((((.(((((((..........(((((....)))))...))))))).)).................(((......)))))))..)))... ( -21.52) >DroYak_CAF1 12763 98 + 1 UUUUGAUUUUUAUGUCCCGACGGAUCCUAACUUAAG-UUUAUUUCUUAUGCCGUCGGAACAUACAAAUUUUU--UUUUUAUAUAUUCGAGUUGCUUUUACU ....(((((.(((((.(((((((.........((((-.......))))..))))))).))))).)))))...--........................... ( -19.30) >consensus UUUUGAUUUCCAUGUCCCGACGGAGCUUAACUUAAAAAUGAUUCCUGUUGCCGUCGGAACUUAAAUAU_____AUUUUCAUAUAUUUCAGUAGCUUUUACU .............((.(((((((...........................))))))).))......................................... (-11.49 = -11.69 + 0.20)

| Location | 8,401,662 – 8,401,754 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 75.50 |
| Mean single sequence MFE | -23.05 |
| Consensus MFE | -14.89 |
| Energy contribution | -15.09 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -3.68 |
| Structure conservation index | 0.65 |
| SVM decision value | 4.29 |
| SVM RNA-class probability | 0.999862 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
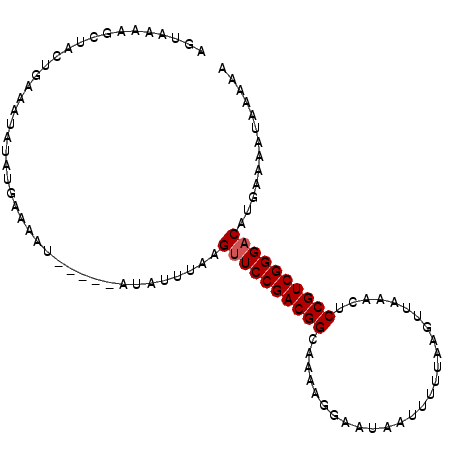
>2R_DroMel_CAF1 8401662 92 - 20766785 AGUAAAAGCUAUUGAAA----UGAAAAU-----AUAGCUAAGGUCCGACGGCAAUAGGAAUCAUUUUUAAAUAAACCUCCGUCGGGACAUGAAAACACAAA ......((((((.....----.......-----))))))..(..(((((((....(((.................))))))))))..)............. ( -19.33) >DroSec_CAF1 12632 96 - 1 AAUCAAAGCUACUGAAAUAUAUGAAAAU-----AUAUUUAAGUUCCGACGGCAACAGGAAUCAUUUUUAAGUUAAACUCCGUCGGGACAUGGGAAUAACAA ........(((...((((((((....))-----))))))..((((((((((.(((..(((.....)))..))).....)))))))))).)))......... ( -23.90) >DroSim_CAF1 12577 96 - 1 AGUAAAAGCUACUGAAAUAUAUGAAAAU-----AUAUUUAAGUUCCGACGGCAACAGGAAUUAUUUUUAAAUUAAACUCCGUCGGGACAUGCGAAUUACAA .((((..((.....((((((((....))-----))))))..((((((((((......((.(((....))).)).....))))))))))..))...)))).. ( -27.50) >DroEre_CAF1 14809 101 - 1 AGAGAGAGCAACUCAAAUAUAUGAAAAUUAAAAUUUUAUAAGUUCCGACGGCAAAAGAAAUAAUUCUUAAGUUAGGAUCCGUCGGGACAUCGAACUCAAAG ...(((..(..........(((((((.......))))))).((((((((((...........(((((......)))))))))))))))...)..))).... ( -24.00) >DroYak_CAF1 12763 98 - 1 AGUAAAAGCAACUCGAAUAUAUAAAAA--AAAAAUUUGUAUGUUCCGACGGCAUAAGAAAUAAA-CUUAAGUUAGGAUCCGUCGGGACAUAAAAAUCAAAA ...........................--....((((.(((((((((((((.((....(((...-.....)))...))))))))))))))).))))..... ( -20.50) >consensus AGUAAAAGCUACUGAAAUAUAUGAAAAU_____AUAUUUAAGUUCCGACGGCAAAAGGAAUAAUUUUUAAGUUAAACUCCGUCGGGACAUGAAAAUAAAAA .........................................((((((((((...........................))))))))))............. (-14.89 = -15.09 + 0.20)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:25:32 2006