
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,287,988 – 8,288,086 |
| Length | 98 |
| Max. P | 0.914389 |

| Location | 8,287,988 – 8,288,086 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 91.24 |
| Mean single sequence MFE | -37.42 |
| Consensus MFE | -29.32 |
| Energy contribution | -29.16 |
| Covariance contribution | -0.16 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.78 |
| SVM decision value | 1.10 |
| SVM RNA-class probability | 0.914389 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8287988 98 + 20766785 -ACCGACCUGG---CAGCUGACUGACUGUCGGGAAUGCGAAAUCCAGAAUUCAGAGUCCUGGUGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCG -.....(((((---((((.....).))))))))((((((....((((.((.....)).)))).((((((.(.......)))))))))))))(((....))). ( -35.30) >DroSec_CAF1 13484 101 + 1 -AACGACCUGGCGGCAGCUGACUGACUGUCGGGAAUGCGGAAUCCAGAAUUCAGAGUCCUGGUGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCG -.....(((((((((((....))).))))))))((((((....((((.((.....)).)))).((((((.(.......)))))))))))))(((....))). ( -38.10) >DroSim_CAF1 9334 101 + 1 -AACGACCUGGCGGCAGCUGACUGACUGUCGGGAAUGCGGAAUCCAGAAUUCAGAGUCCUGGUGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCG -.....(((((((((((....))).))))))))((((((....((((.((.....)).)))).((((((.(.......)))))))))))))(((....))). ( -38.10) >DroEre_CAF1 9783 95 + 1 UACCAACCUGG---CAGC----UGACUGUCGGGAAUGCGGAAUCCAGAAUUCAGAGUCCUGGAGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCA ......(((((---(((.----...))))))))((((((...(((((.((.....)).)))))((((((.(.......)))))))))))))(((....))). ( -37.50) >DroYak_CAF1 9860 95 + 1 UACCAACCUGG---CAGC----UGACUGUCGGGGAUGCGGAAUCCAGUAUUCAAAGUCCUGGAGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCA ......(((((---(((.----...))))))))((((((...(((((.((.....)).)))))((((((.(.......)))))))))))))(((....))). ( -38.10) >consensus _ACCGACCUGG___CAGCUGACUGACUGUCGGGAAUGCGGAAUCCAGAAUUCAGAGUCCUGGUGCCGAGAGUCGUUAUCCUCGGCCGCAUUGCUUUUUAGCG ......(((((...(((........))))))))((((((....((((.((.....)).)))).((((((.(.......)))))))))))))(((....))). (-29.32 = -29.16 + -0.16)

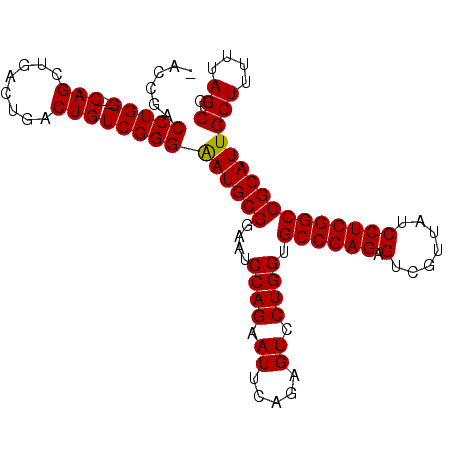
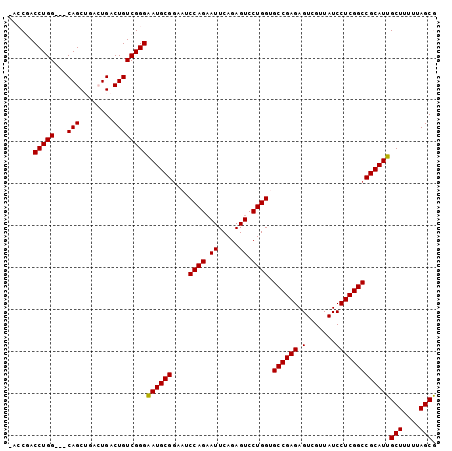
| Location | 8,287,988 – 8,288,086 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 91.24 |
| Mean single sequence MFE | -33.36 |
| Consensus MFE | -28.86 |
| Energy contribution | -29.82 |
| Covariance contribution | 0.96 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.09 |
| SVM RNA-class probability | 0.913416 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8287988 98 - 20766785 CGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCACCAGGACUCUGAAUUCUGGAUUUCGCAUUCCCGACAGUCAGUCAGCUG---CCAGGUCGGU- .(((....)))(((((((((((((........))))))).((((((.......))))))....)))))).(((((...(((....)))---....))))).- ( -37.20) >DroSec_CAF1 13484 101 - 1 CGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCACCAGGACUCUGAAUUCUGGAUUCCGCAUUCCCGACAGUCAGUCAGCUGCCGCCAGGUCGUU- .(((....)))(((((((((((((........))))))).((((((.......))))))....))))))...(((.....)))(((.(((....))).)))- ( -33.70) >DroSim_CAF1 9334 101 - 1 CGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCACCAGGACUCUGAAUUCUGGAUUCCGCAUUCCCGACAGUCAGUCAGCUGCCGCCAGGUCGUU- .(((....)))(((((((((((((........))))))).((((((.......))))))....))))))...(((.....)))(((.(((....))).)))- ( -33.70) >DroEre_CAF1 9783 95 - 1 UGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCUCCAGGACUCUGAAUUCUGGAUUCCGCAUUCCCGACAGUCA----GCUG---CCAGGUUGGUA ((((((.....(((((((((((((........)))))))(((((((.......)))))))...))))))((.(.(((...----.)))---.).)))))))) ( -32.00) >DroYak_CAF1 9860 95 - 1 UGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCUCCAGGACUUUGAAUACUGGAUUCCGCAUCCCCGACAGUCA----GCUG---CCAGGUUGGUA ((((....))))((((((((((((........)))))))(((((...........)))))...)))))..(((((.((..----...)---)...))))).. ( -30.20) >consensus CGCUAAAAAGCAAUGCGGCCGAGGAUAACGACUCUCGGCACCAGGACUCUGAAUUCUGGAUUCCGCAUUCCCGACAGUCAGUCAGCUG___CCAGGUCGGU_ .(((....)))(((((((((((((........))))))).((((((.......))))))....)))))).(((((...(((....))).......))))).. (-28.86 = -29.82 + 0.96)

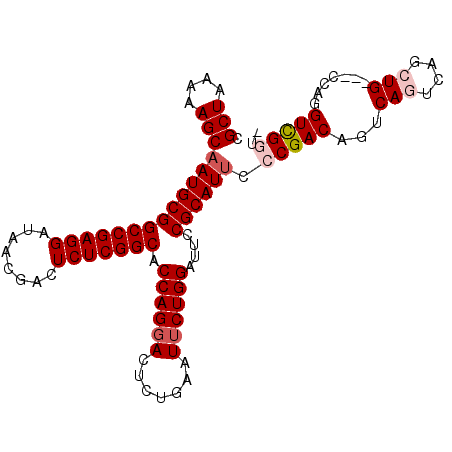
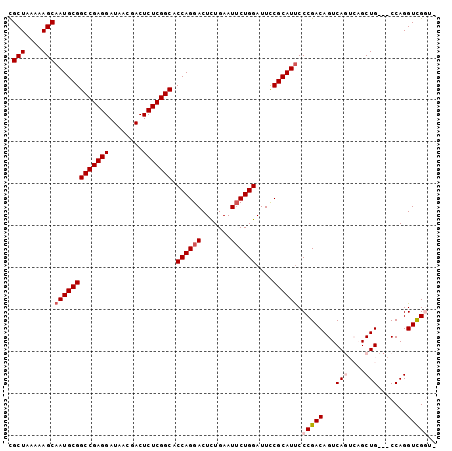
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:51 2006