
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,277,386 – 8,277,543 |
| Length | 157 |
| Max. P | 0.974973 |

| Location | 8,277,386 – 8,277,503 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 92.45 |
| Mean single sequence MFE | -40.15 |
| Consensus MFE | -33.88 |
| Energy contribution | -35.00 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.85 |
| Structure conservation index | 0.84 |
| SVM decision value | 1.73 |
| SVM RNA-class probability | 0.974243 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
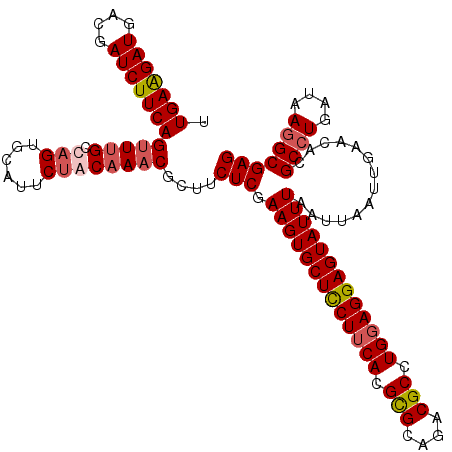
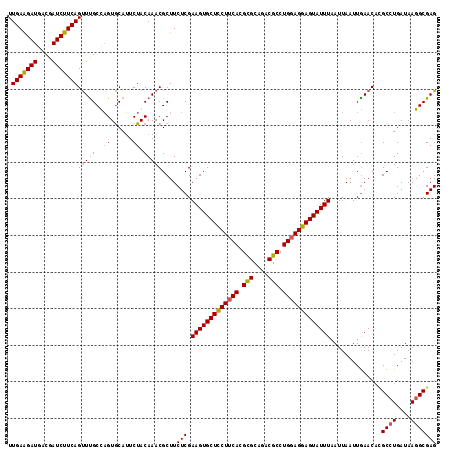
>2R_DroMel_CAF1 8277386 117 + 20766785 UUGAAGAUGACGAUCUUCAGUUUGCCAGUGCAUUCUACAAACGCUUCUCGAAGUGCUCCUUCACGUGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACAUGCCUGAUAAGGCGAG .(((((((....)))))))(((((..((......)).)))))....(((.(((((((((((((.(((....))).)))))))))))))..............((((....))))))) ( -38.70) >DroSim_CAF1 3736 117 + 1 UUGAAGAUGACGAUCUUCAGUUCGCCCGUACAGUCUACAAACGCUUCUCGAAGUGCUCCUUCACGCGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACUAGCGUGAUAAGGCGAG ((((((((....)))))))).(((((.(((.....)))..(((((..((((((((((((((((.(((....))).)))))))))))).......))))...))))).....))))). ( -42.91) >DroEre_CAF1 4004 117 + 1 UUGAGGAUGGCGAUCUUCAGCUUGCUAGUGCAUUCUACAAACGCUUCUCGAAGUGCUUCUUCACGCGCAGACGCCUGGAGGAGUAUUUCAUUAAUUAAACACGCCUGAUAAGGCGAG ..((((..((((......((..(((....)))..)).....))))))))((((((((((((((.(((....))).))))))))))))))............(((((....))))).. ( -42.70) >DroYak_CAF1 3718 117 + 1 UUGAAGAUGGCGAUCUUCAGUUUGCUAGUGCAUUCUACAAACGCUUCUCGAAGUGCUUCUUCACGCGCAGACGCCUGUAGGAGUAUUUAAUUAAUUGAACACGCCUGAUAAGGCGAG .(((((((....)))))))(((((.(((......))))))))........((((((((((.((.(((....))).)).)))))))))).............(((((....))))).. ( -36.30) >consensus UUGAAGAUGACGAUCUUCAGUUUGCCAGUGCAUUCUACAAACGCUUCUCGAAGUGCUCCUUCACGCGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACACGCCUGAUAAGGCGAG .(((((((....)))))))(((((.(((......))))))))....(((.(((((((((((((.(((....))).)))))))))))))..............((((....))))))) (-33.88 = -35.00 + 1.12)

| Location | 8,277,426 – 8,277,543 |
|---|---|
| Length | 117 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 93.30 |
| Mean single sequence MFE | -36.18 |
| Consensus MFE | -32.74 |
| Energy contribution | -32.43 |
| Covariance contribution | -0.31 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.90 |
| SVM decision value | 1.74 |
| SVM RNA-class probability | 0.974973 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
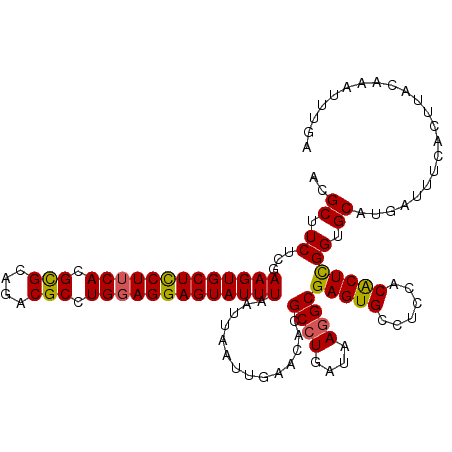
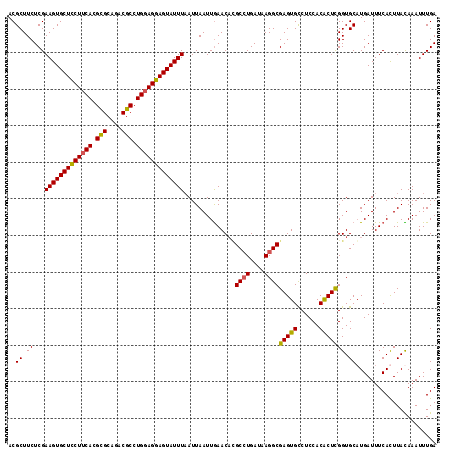
>2R_DroMel_CAF1 8277426 117 + 20766785 ACGCUUCUCGAAGUGCUCCUUCACGUGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACAUGCCUGAUAAGGCGAGUGCCUCCACACUCGGUGCAUGAUUUCACUUACAAAUUUGA ..........(((((((((((((.(((....))).)))))))))))))...(((.(((((((((((....)).((((((......))))))..)))))..)))).)))......... ( -35.70) >DroSim_CAF1 3776 117 + 1 ACGCUUCUCGAAGUGCUCCUUCACGCGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACUAGCGUGAUAAGGCGAGUGCCUCCACACUCGGUGCUUGAUUUCACUUACAAAUUUGA ..(((..((((((((((((((((.(((....))).)))))))))))).......))))...)))((((.....((((..((.........))..))))...))))............ ( -38.51) >DroEre_CAF1 4044 117 + 1 ACGCUUCUCGAAGUGCUUCUUCACGCGCAGACGCCUGGAGGAGUAUUUCAUUAAUUAAACACGCCUGAUAAGGCGAGUGCCUCCACACUCGGUGCAUGAUUUCAUUUAUAAAUUUGA .........((((((((((((((.(((....))).))))))))))))))...(((((..(((((((....))))(((((......))))).)))..)))))................ ( -39.60) >DroYak_CAF1 3758 117 + 1 ACGCUUCUCGAAGUGCUUCUUCACGCGCAGACGCCUGUAGGAGUAUUUAAUUAAUUGAACACGCCUGAUAAGGCGAGUGUCUCCACGCUUGGUGCAUGAUUUCAUUUAUAAAUUUGA .(((((....))))).....(((.((((...(((((.((((.((.(((((....))))).)).))))...)))))(((((....)))))..)))).))).................. ( -30.90) >consensus ACGCUUCUCGAAGUGCUCCUUCACGCGCAGACGCCUGGAGGAGUAUUUAAUUAAUUGAACACGCCUGAUAAGGCGAGUGCCUCCACACUCGGUGCAUGAUUUCACUUACAAAUUUGA ..((.((...(((((((((((((.(((....))).)))))))))))))..............((((....))))(((((......))))))).))...................... (-32.74 = -32.43 + -0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:24:41 2006