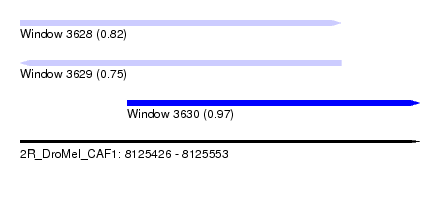
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,125,426 – 8,125,553 |
| Length | 127 |
| Max. P | 0.966621 |

| Location | 8,125,426 – 8,125,528 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 98.53 |
| Mean single sequence MFE | -35.23 |
| Consensus MFE | -33.00 |
| Energy contribution | -33.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.16 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.815355 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8125426 102 + 20766785 AGGAAGGUGGCAAACAGCCCAAGGAACUGCCAUCUACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACA .((.((((((((.....((...))...)))))))).))...(((.........((((((((....))))))))...(.((((.....)))).).)))..... ( -35.70) >DroSec_CAF1 11725 102 + 1 AGGACGGUGGCAAACAGCCCAAGGAACUGCCAUCCACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACA ....(((((((.....)))...((((.((((((........((.......)).((((((((....)))))))).....)))))).))))))))......... ( -33.90) >DroSim_CAF1 2589 102 + 1 AGGAAGGUGGCAAACAGCCCAAGGAACUGCCAUCCACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACA .((..(((((((.....((...))...)))))))..))...(((.........((((((((....))))))))...(.((((.....)))).).)))..... ( -33.40) >DroYak_CAF1 12123 102 + 1 AGGAAGGUGGCAAACAGCCCAAGGAGCUGCCAUCCACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACA .(((.(((((....(((((....).))))....)))))...)))..(((....((((((((....)))))))).............(((.....)))))).. ( -37.90) >consensus AGGAAGGUGGCAAACAGCCCAAGGAACUGCCAUCCACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACA .((..(((((((.....((...))...)))))))..))...(((.........((((((((....))))))))...(.((((.....)))).).)))..... (-33.00 = -33.00 + 0.00)

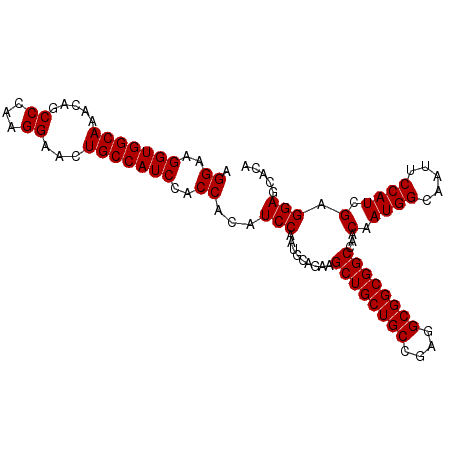
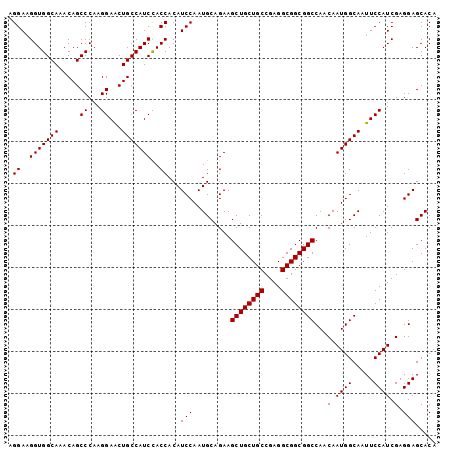
| Location | 8,125,426 – 8,125,528 |
|---|---|
| Length | 102 |
| Sequences | 4 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 98.53 |
| Mean single sequence MFE | -40.23 |
| Consensus MFE | -38.62 |
| Energy contribution | -38.88 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.94 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.753221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8125426 102 - 20766785 UGUGCUCCUCGAUGGAAUUGCCAUUGUUGGCCGCCGCCUCGGCAGCAGCUUCUGCAUUGGAUGUGGUAGAUGGCAGUUCCUUGGGCUGUUUGCCACCUUCCU .........(((.((((((((((((....(((((..((...((((......))))...))..))))).)))))))))))))))(((.....)))........ ( -41.10) >DroSec_CAF1 11725 102 - 1 UGUGCUCCUCGAUGGAAUUGCCAUUGUUGGCCGCCGCCUCGGCAGCAGCUUCUGCAUUGGAUGUGGUGGAUGGCAGUUCCUUGGGCUGUUUGCCACCGUCCU .........(((.(((((((((((......(((((((.((.(..((((...))))..).)).)))))))))))))))))))))(((.....)))........ ( -39.20) >DroSim_CAF1 2589 102 - 1 UGUGCUCCUCGAUGGAAUUGCCAUUGUUGGCCGCCGCCUCGGCAGCAGCUUCUGCAUUGGAUGUGGUGGAUGGCAGUUCCUUGGGCUGUUUGCCACCUUCCU .........(((.(((((((((((......(((((((.((.(..((((...))))..).)).)))))))))))))))))))))(((.....)))........ ( -39.20) >DroYak_CAF1 12123 102 - 1 UGUGCUCCUCGAUGGAAUUGCCAUUGUUGGCCGCCGCCUCGGCAGCAGCUUCUGCAUUGGAUGUGGUGGAUGGCAGCUCCUUGGGCUGUUUGCCACCUUCCU .(.(((...((((((.....))))))..))))((((...)))).((((...))))...(((...(((((..(((((((.....)))))))..))))).))). ( -41.40) >consensus UGUGCUCCUCGAUGGAAUUGCCAUUGUUGGCCGCCGCCUCGGCAGCAGCUUCUGCAUUGGAUGUGGUGGAUGGCAGUUCCUUGGGCUGUUUGCCACCUUCCU .........(((.((((((((((((....(((((..((...((((......))))...))..))))).)))))))))))))))(((.....)))........ (-38.62 = -38.88 + 0.25)
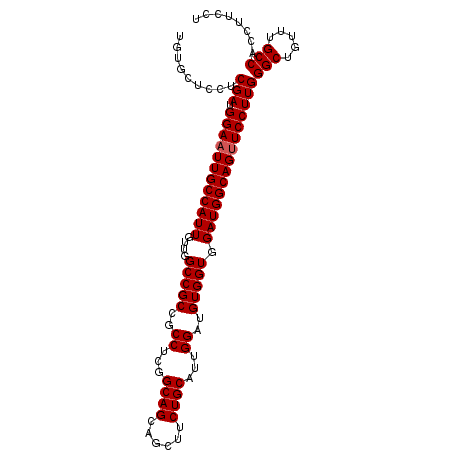


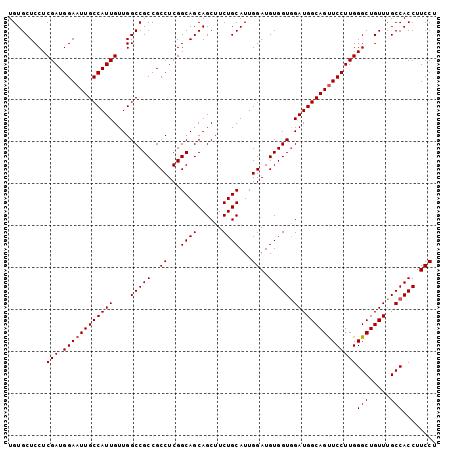
| Location | 8,125,460 – 8,125,553 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 93 |
| Reading direction | forward |
| Mean pairwise identity | 98.49 |
| Mean single sequence MFE | -26.00 |
| Consensus MFE | -25.48 |
| Energy contribution | -25.48 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.60 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.60 |
| SVM RNA-class probability | 0.966621 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 8125460 93 + 20766785 UACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACAAGCAGAAGAAACUAGGUAAUGACAC ((((...(((.........((((((((....))))))))...(.((((.....)))).).)))((....))...........))))....... ( -26.60) >DroSec_CAF1 11759 93 + 1 CACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACAAGCAGAAGAAACUAGGUAAUGACAC .(((...(((.........((((((((....))))))))...(.((((.....)))).).)))((....))...........)))........ ( -25.50) >DroSim_CAF1 2623 93 + 1 CACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACAAGCAGAAGAAACUAGGUAAUGACAC .(((...(((.........((((((((....))))))))...(.((((.....)))).).)))((....))...........)))........ ( -25.50) >DroEre_CAF1 2761 93 + 1 CACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAACUCCAUCGAGGAGCACAAGCAGAAGAAAUUAGGUAAUGACAC .(((........(((....((((((((....))))))))......((((..(((....)))..)).)).))).((....)).)))........ ( -26.90) >DroYak_CAF1 12157 93 + 1 CACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACAAGCAGAAGAAAUUAGGUAAUGACAC .(((...(((.........((((((((....))))))))...(.((((.....)))).).)))((....))...........)))........ ( -25.50) >consensus CACCACAUCCAAUGCAGAAGCUGCUGCCGAGGCGGCGGCCAACAAUGGCAAUUCCAUCGAGGAGCACAAGCAGAAGAAACUAGGUAAUGACAC .(((...(((.........((((((((....))))))))...(.((((.....)))).).)))((....))...........)))........ (-25.48 = -25.48 + 0.00)

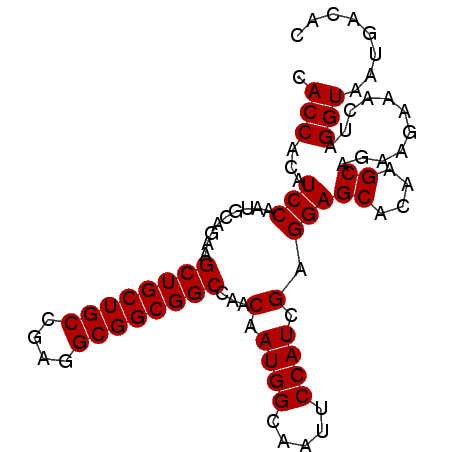
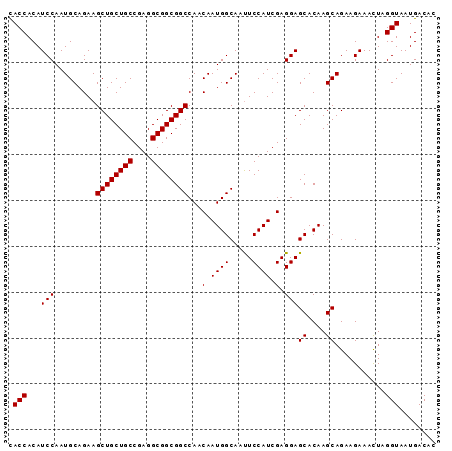
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:23:42 2006