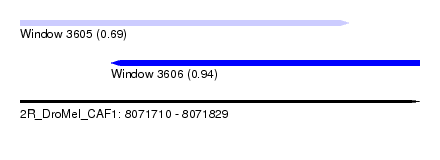

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 8,071,710 – 8,071,829 |
| Length | 119 |
| Max. P | 0.944507 |

| Location | 8,071,710 – 8,071,808 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 98 |
| Reading direction | forward |
| Mean pairwise identity | 72.90 |
| Mean single sequence MFE | -24.69 |
| Consensus MFE | -8.26 |
| Energy contribution | -9.62 |
| Covariance contribution | 1.36 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.33 |
| SVM decision value | 0.32 |
| SVM RNA-class probability | 0.687117 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
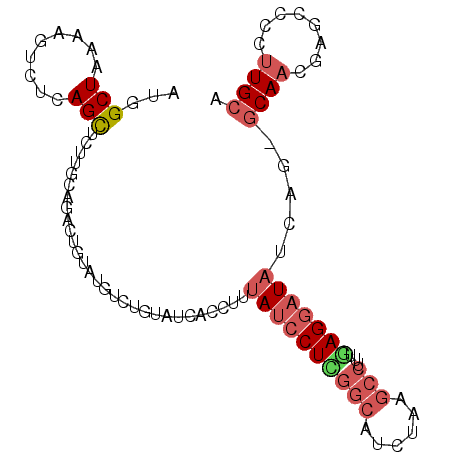
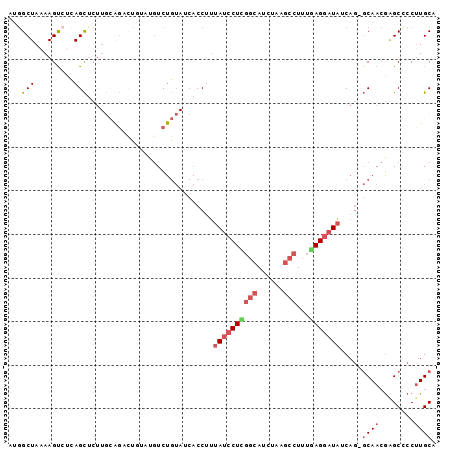
>2R_DroMel_CAF1 8071710 98 + 20766785 AUGGCUAAAAGUCUCAGCACUUGAAGACCGUCUGUCUGUAUCACCUUUAUCCUUGGCAUCUAAGCCUUUGAGGAUAUCAGGGCAACAGGCCCCCUGCA ((((((..((((......))))..)).))))..((((((....(((.((((((((((......)))...)))))))..)))...))))))........ ( -27.30) >DroSec_CAF1 36713 97 + 1 AUGGCUAAAAGUCUCAGCUCCUGCAGAUUUUCUGUCUGUAUCACCUUUAUCCUCGGCAUCUAAGCCUUUAAGGAUAUCAU-GCAACGCGCCCCUUGCA ..(((..((((((((((...))).)))))))..)))(((((......((((((.(((......)))....))))))..))-)))..(((.....))). ( -22.40) >DroEre_CAF1 35194 97 + 1 AUGGCUAAAAGCCUCAGCUCUUCCGGCCUGUAUGUCUGUAUCACCUUUAUCCUCGGCAUCUAAGCCUUUGAGGAUAUCAG-GCAACGAGCACCUUGCG ..(((.....)))...((((.....(((((.................((((((((((......)))...))))))).)))-))...))))........ ( -29.27) >DroYak_CAF1 35117 93 + 1 AUAGCUAAAAGUACUAGCUCUUGCAGCCUG----UCUGUAUCACCUUUAUCCUCGGCAUCUAAGCCUUUGAGGAUAUCAG-GCAAGGAGCCCCUUGCG ..(((((.......)))))..(((((....----.)))))....((.((((((((((......)))...)))))))..))-((((((....)))))). ( -28.30) >DroAna_CAF1 35700 78 + 1 A-GGCUAGAAGUAGAAGU-CU---------CAUGUC-------CCAGUAUCCUUC-UAGCCAAUCCUUUAAGCCUCACAG-CCAACGAGCUCCUUGCA .-(((((((((..((...-((---------......-------..))..))))))-)))))..........((.....((-(......)))....)). ( -16.20) >consensus AUGGCUAAAAGUCUCAGCUCUUGCAGACUGUAUGUCUGUAUCACCUUUAUCCUCGGCAUCUAAGCCUUUGAGGAUAUCAG_GCAACGAGCCCCUUGCA ...(((.........))).............................((((((((((......)))...))))))).....((((........)))). ( -8.26 = -9.62 + 1.36)

| Location | 8,071,737 – 8,071,829 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 77.90 |
| Mean single sequence MFE | -25.06 |
| Consensus MFE | -12.74 |
| Energy contribution | -13.74 |
| Covariance contribution | 1.00 |
| Combinations/Pair | 1.42 |
| Mean z-score | -2.30 |
| Structure conservation index | 0.51 |
| SVM decision value | 1.34 |
| SVM RNA-class probability | 0.944507 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
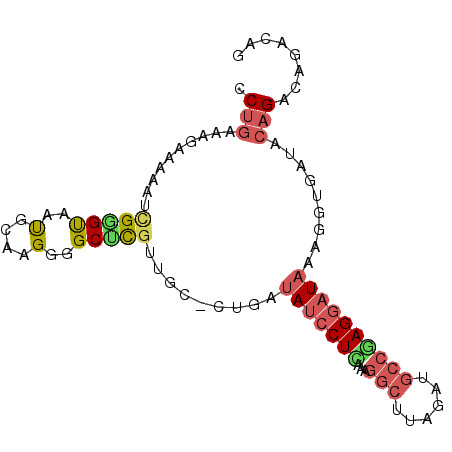
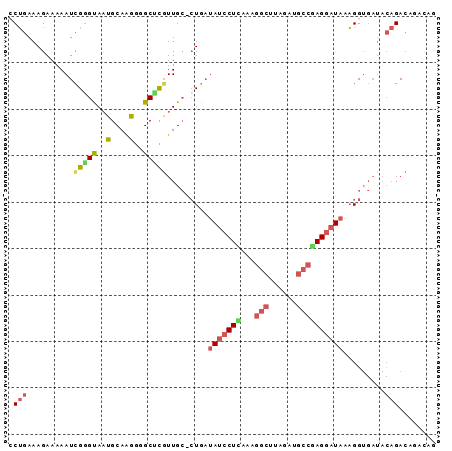
>2R_DroMel_CAF1 8071737 92 - 20766785 CCUGAAAGAAAAAUCGGGUAAUGCAGGGGGCCUGUUGCCCUGAUAUCCUCAAAGGCUUAGAUGCCAAGGAUAAAGGUGAUACAGACAGACGG .(((........((((((((..(((((...))))))))))...((((((....(((......))).))))))..)))........))).... ( -25.39) >DroSec_CAF1 36740 91 - 1 CCUGAAAGAAAAAUCGGGCAAUGCAAGGGGCGCGUUGC-AUGAUAUCCUUAAAGGCUUAGAUGCCGAGGAUAAAGGUGAUACAGACAGAAAA .(((.........(((.(((((((.......)))))))-.)))(((((((...(((......)))))))))).........)))........ ( -25.60) >DroEre_CAF1 35221 91 - 1 GCUGAAAGAAAAAUCGGGUAACGCAAGGUGCUCGUUGC-CUGAUAUCCUCAAAGGCUUAGAUGCCGAGGAUAAAGGUGAUACAGACAUACAG .(((...........(((((.(....).)))))((..(-((..(((((((...(((......)))))))))).)))..)).)))........ ( -31.50) >DroYak_CAF1 35144 87 - 1 GCUGAAAGAAAAAUCGGGUAACGCAAGGGGCUCCUUGC-CUGAUAUCCUCAAAGGCUUAGAUGCCGAGGAUAAAGGUGAUACAGA----CAG .(((...........((((..(....)..)))).(..(-((..(((((((...(((......)))))))))).)))..)..))).----... ( -29.90) >DroAna_CAF1 35719 80 - 1 CCAGAAAGAAAAAUUGAGUAUUGCAAGGAGCUCGUUGG-CUGUGAGGCUUAAAGGAUUGGCUA-GAAGGAUACUGG-------GACAUG--- ((((.........((((((.(..((....(((....))-)))..).))))))((......)).-........))))-------......--- ( -12.90) >consensus CCUGAAAGAAAAAUCGGGUAAUGCAAGGGGCUCGUUGC_CUGAUAUCCUCAAAGGCUUAGAUGCCGAGGAUAAAGGUGAUACAGACAGACAG .(((..........(((((..(....)..))))).........(((((((...(((......)))))))))).........)))........ (-12.74 = -13.74 + 1.00)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:23:19 2006