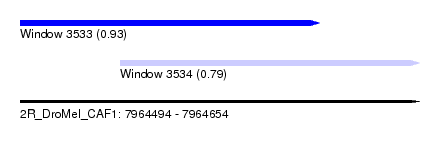

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,964,494 – 7,964,654 |
| Length | 160 |
| Max. P | 0.930339 |

| Location | 7,964,494 – 7,964,614 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.89 |
| Mean single sequence MFE | -35.98 |
| Consensus MFE | -35.20 |
| Energy contribution | -34.82 |
| Covariance contribution | -0.39 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.12 |
| Structure conservation index | 0.98 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.930339 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
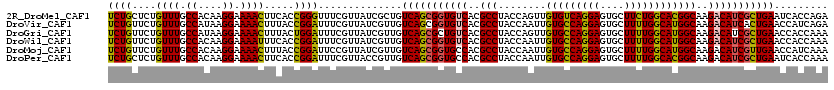
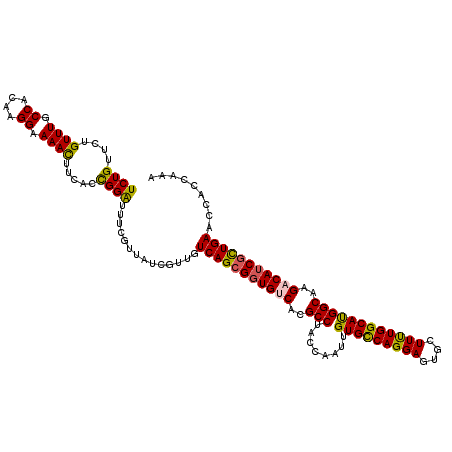
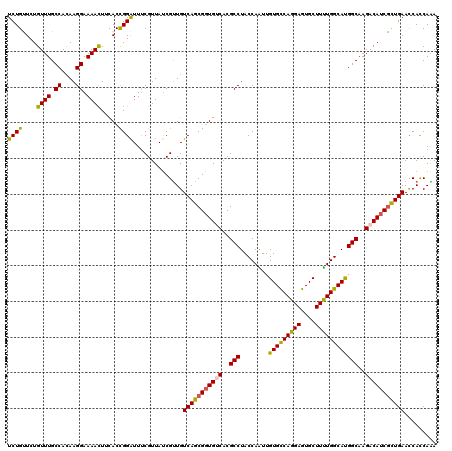
>2R_DroMel_CAF1 7964494 120 + 20766785 UCUGCUCUGUUUGCCACAAGGAAAACUUCACCGGGUUUCGUUAUCGCUGUCAGCGGUGUCACGCCUACCAGUUGUGUCAGGAGUGCUUCUGGCACGGCAAGACAUCGCUGAAUCACCAGA .....((((...((...((.((((.((......)))))).))...))..(((((((((((..(.....).((((((((((((....))))))))))))..))))))))))).....)))) ( -40.30) >DroVir_CAF1 818 120 + 1 UCUGUUCUGUUUGCCAUAAGGAAAACUUUACCGGAUUUCGUUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCACUGAACCAUCAGA .....((((...((.((((.((((.((.....)).)))).)))).))..((((.((((((..(((........(((((((((....))))))))))))..)))))).)))).....)))) ( -34.90) >DroGri_CAF1 423 120 + 1 UCUGUUCUGUUUGCCAUAAGGAAAACUUUACUGGAUUUCGUUAUCGUUGUCAGCGCUGUCACGCCUACCAGUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAA ...((...((((.((....)).))))...))(((....((....))...((((((.((((..(((........(((((((((....))))))))))))..)))).))))))....))).. ( -34.70) >DroWil_CAF1 19306 120 + 1 UCUGUUCUGUUUGCCACAAGGAAAAUUUCACCGGAUUUCGUUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAA ((((....((((.((....)).)))).....))))..........((..(((((((((((..(((........(((((((((....))))))))))))..)))))))))))...)).... ( -37.30) >DroMoj_CAF1 856 120 + 1 UCUGUUCUGUUUGCCACAAGGAAAACUUUACCGGAUUCCGUUAUCGUUGUCAGCGGUGCCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGUUGAACCAUCAAA ...((((((((((((...(((.......((((((((..((....))..)))..))))).....))).......(((((((((....)))))))))))).))))).....))))....... ( -31.60) >DroPer_CAF1 14193 120 + 1 UCUGCUCUGUUUGCCACAAGGAAAACUUCACCGGAUUUCGUUACCGUUGUCAGCGGUGCCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCACGGCAAGACAUCGCUGAAUCACCAAA ((.((..((((((((...(((.......((((((((..((....))..)))..))))).....))).......(((((((((....)))))))))))).)))))..)).))......... ( -37.10) >consensus UCUGUUCUGUUUGCCACAAGGAAAACUUCACCGGAUUUCGUUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAA ((((....((((.((....)).)))).....))))..............(((((((((((..(((........(((((((((....))))))))))))..)))))))))))......... (-35.20 = -34.82 + -0.39)
| Location | 7,964,534 – 7,964,654 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 93.72 |
| Mean single sequence MFE | -38.05 |
| Consensus MFE | -35.89 |
| Energy contribution | -36.25 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.07 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.59 |
| SVM RNA-class probability | 0.791634 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
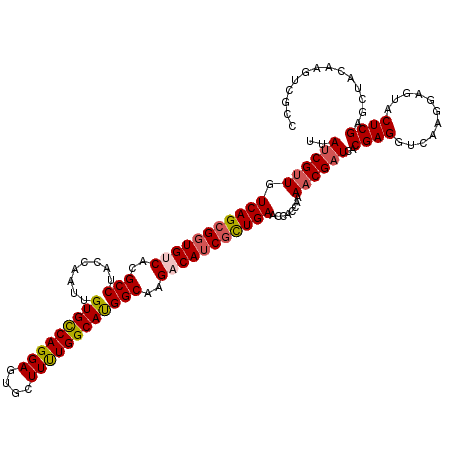
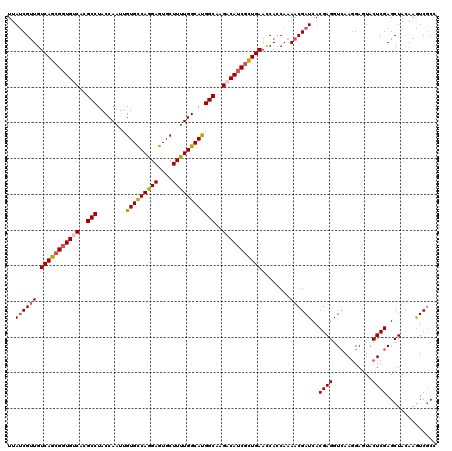
>2R_DroMel_CAF1 7964534 120 + 20766785 UUAUCGCUGUCAGCGGUGUCACGCCUACCAGUUGUGUCAGGAGUGCUUCUGGCACGGCAAGACAUCGCUGAAUCACCAGAACGAUCACGAGGUCAAGGAGUACUCGAGCUACAAGUCGCC ......((((((((((((((..(.....).((((((((((((....))))))))))))..))))))))))).....)))..((((....((.((.((.....)).)).))....)))).. ( -42.40) >DroVir_CAF1 858 120 + 1 UUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCACUGAACCAUCAGAACGAUCACGAGGUCAAGGAGUACUCGAGCUACAAGUCGCC ...((..((((((.((((((..(((........(((((((((....))))))))))))..)))))).))))..))...)).((((....((.((.((.....)).)).))....)))).. ( -35.20) >DroGri_CAF1 463 120 + 1 UUAUCGUUGUCAGCGCUGUCACGCCUACCAGUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAAACGAUCACGAGGUCAAGGAGUACUCGAGCUACAAGUCCCC ..((((((.((((((.((((..(((........(((((((((....))))))))))))..)))).))))))........))))))..((((...........)))).............. ( -36.70) >DroWil_CAF1 19346 120 + 1 UUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAAACGAUCACGAGGUCAAGGAAUACUCGAGCUACAAAUCGCC ..((((((.(((((((((((..(((........(((((((((....))))))))))))..)))))))))))........))))))..((((...........)))).((........)). ( -40.90) >DroMoj_CAF1 896 120 + 1 UUAUCGUUGUCAGCGGUGCCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGUUGAACCAUCAAAACGAUCACGAGGUGAAAGAGUACUCGAGCUAUAAGUCGCC ..((((((.(((((((((.(..(((........(((((((((....))))))))))))..).)))))))))........)))))).....(((((...(((......))).....))))) ( -36.00) >DroPer_CAF1 14233 120 + 1 UUACCGUUGUCAGCGGUGCCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCACGGCAAGACAUCGCUGAAUCACCAAAACGAUCACGAGGUCAAGGAGUACUCGAGCUACAAGUCGCC .....(...(((((((((.(..(((........(((((((((....))))))))))))..).)))))))))..).......((((....((.((.((.....)).)).))....)))).. ( -37.10) >consensus UUAUCGUUGUCAGCGGUGUCACGCCUACCAAUUGUGCCAGGAGUGCUUUUGGCAUGGCAAGACAUCGCUGAACCACCAAAACGAUCACGAGGUCAAGGAGUACUCGAGCUACAAGUCGCC ..((((((.(((((((((((..(((........(((((((((....))))))))))))..)))))))))))........))))))..((((...........)))).............. (-35.89 = -36.25 + 0.36)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:22:10 2006