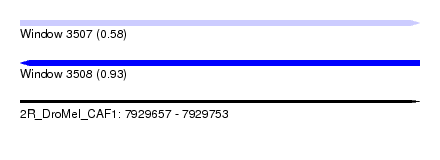

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,929,657 – 7,929,753 |
| Length | 96 |
| Max. P | 0.932739 |

| Location | 7,929,657 – 7,929,753 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | forward |
| Mean pairwise identity | 91.61 |
| Mean single sequence MFE | -16.96 |
| Consensus MFE | -12.68 |
| Energy contribution | -12.88 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.09 |
| SVM RNA-class probability | 0.577296 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
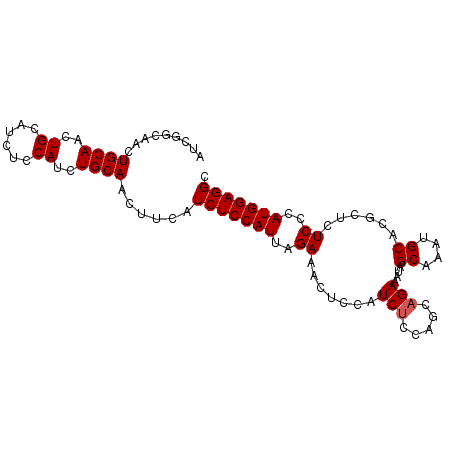
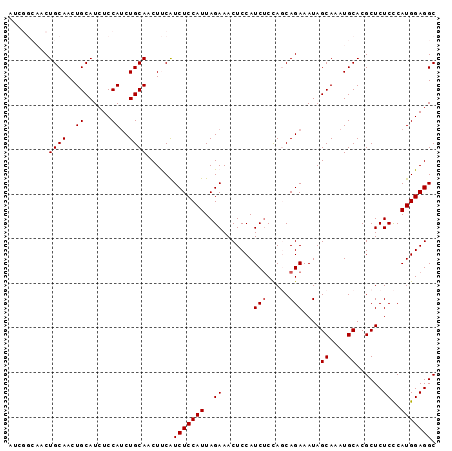
>2R_DroMel_CAF1 7929657 96 + 20766785 AUCGGCAACUGCAACUGCAUAUCCAUCUGCAACUUCAUCUCCAUUAGAAACUCCAUCACCAGCAGAAAUAGCAAAUGCACGCUCUCCCAUGGAGGC ...(....)((((..(((.(((...(((((...............................))))).))))))..)))).((.((((...)))))) ( -17.08) >DroSec_CAF1 3757 96 + 1 AUCGGCAACUGCAACUGCAUCUCCAUCUGCAACUUCAUCUCCAUUAGAAACUCCAUCUCCAGCAGAAAUAGCAAAUGCACGCUCUCCCAUGGAGGC ...(((..((((...((((........))))......(((.....))).............)))).....((....))..)))((((...)))).. ( -16.50) >DroSim_CAF1 3746 96 + 1 AUCGGCAACUGCAACUGCAUCUCCAUCUGCAACUUCAUCUCCAUUAGAAACUCCAUCUCCAGCAGAAAUAGCAAAUGCACGCUCUCCCAUGGAGGC ...(((..((((...((((........))))......(((.....))).............)))).....((....))..)))((((...)))).. ( -16.50) >DroEre_CAF1 3620 90 + 1 AC------CUGCAACUGCAUCUCCAGCUGCAACUCCAUCUCCAUUAGAAACUCCAUCUCUAUCAGAAAUAGCAAAUGCACGCUCUCUCAUGGAGGC ..------.((((.(((......))).))))......(((((((.(((.......(((.....)))....((....))......))).))))))). ( -18.90) >DroYak_CAF1 3712 90 + 1 AC------CUGCAACUGCAUCUCCAUCUGCAACUCCGUCUCCAUUAGAAACUCCAUCUCUAUCAGAAAUAGCAAAUGCACGCUCUCCCAUGGAGGC ..------.((((..((......))..)))).....((((((((..((.......(((.....)))....((....))......))..)))))))) ( -15.80) >consensus AUCGGCAACUGCAACUGCAUCUCCAUCUGCAACUUCAUCUCCAUUAGAAACUCCAUCUCCAGCAGAAAUAGCAAAUGCACGCUCUCCCAUGGAGGC .........((((..((......))..))))......(((((((..((.......(((.....)))....((....))......))..))))))). (-12.68 = -12.88 + 0.20)

| Location | 7,929,657 – 7,929,753 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 91.61 |
| Mean single sequence MFE | -28.52 |
| Consensus MFE | -24.98 |
| Energy contribution | -25.02 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.59 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.22 |
| SVM RNA-class probability | 0.932739 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
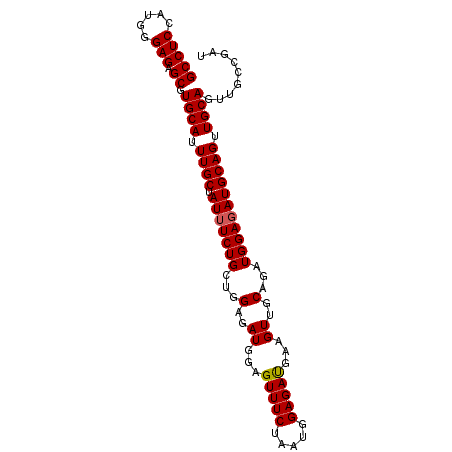
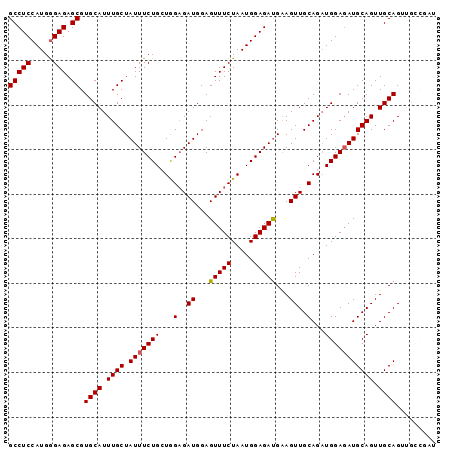
>2R_DroMel_CAF1 7929657 96 - 20766785 GCCUCCAUGGGAGAGCGUGCAUUUGCUAUUUCUGCUGGUGAUGGAGUUUCUAAUGGAGAUGAAGUUGCAGAUGGAUAUGCAGUUGCAGUUGCCGAU ((((((...)))).)).((((.(((((((((((((....(((...(((((.....)))))...))))))))..)))).)))).))))......... ( -26.70) >DroSec_CAF1 3757 96 - 1 GCCUCCAUGGGAGAGCGUGCAUUUGCUAUUUCUGCUGGAGAUGGAGUUUCUAAUGGAGAUGAAGUUGCAGAUGGAGAUGCAGUUGCAGUUGCCGAU ((((((...)))).)).((((.((((.((((((((((..(((...(((((.....)))))...))).))).))))))))))).))))......... ( -29.40) >DroSim_CAF1 3746 96 - 1 GCCUCCAUGGGAGAGCGUGCAUUUGCUAUUUCUGCUGGAGAUGGAGUUUCUAAUGGAGAUGAAGUUGCAGAUGGAGAUGCAGUUGCAGUUGCCGAU ((((((...)))).)).((((.((((.((((((((((..(((...(((((.....)))))...))).))).))))))))))).))))......... ( -29.40) >DroEre_CAF1 3620 90 - 1 GCCUCCAUGAGAGAGCGUGCAUUUGCUAUUUCUGAUAGAGAUGGAGUUUCUAAUGGAGAUGGAGUUGCAGCUGGAGAUGCAGUUGCAG------GU ..((((((((((.((((......)))).))))...(((((((...)))))))))))))......(((((((((......)))))))))------.. ( -30.20) >DroYak_CAF1 3712 90 - 1 GCCUCCAUGGGAGAGCGUGCAUUUGCUAUUUCUGAUAGAGAUGGAGUUUCUAAUGGAGACGGAGUUGCAGAUGGAGAUGCAGUUGCAG------GU ((((((...)))).)).((((.((((.((((((......(((...(((((.....)))))...)))......)))))))))).)))).------.. ( -26.90) >consensus GCCUCCAUGGGAGAGCGUGCAUUUGCUAUUUCUGCUGGAGAUGGAGUUUCUAAUGGAGAUGAAGUUGCAGAUGGAGAUGCAGUUGCAGUUGCCGAU (((((.....))).)).((((.((((.(((((((...(..((...(((((.....)))))...))..)...))))))))))).))))......... (-24.98 = -25.02 + 0.04)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:45 2006