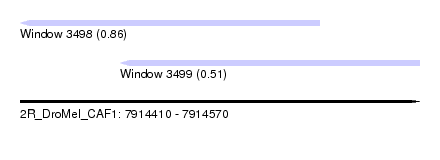

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,914,410 – 7,914,570 |
| Length | 160 |
| Max. P | 0.855023 |

| Location | 7,914,410 – 7,914,530 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 87.83 |
| Mean single sequence MFE | -34.20 |
| Consensus MFE | -25.62 |
| Energy contribution | -25.62 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.23 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.75 |
| SVM decision value | 0.80 |
| SVM RNA-class probability | 0.855023 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
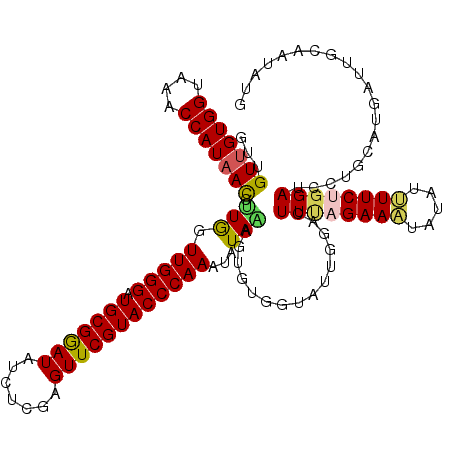
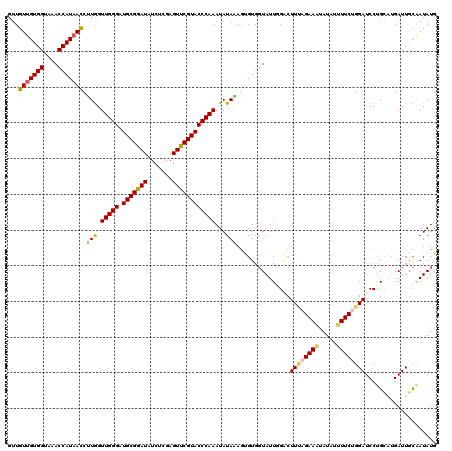
>2R_DroMel_CAF1 7914410 120 - 20766785 GUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUACAAUGUGGUAUUUGACUUUAGAAAUAUAUUUUCUGGAUCCUGCAUGAUUUCAAUAUG ...(((((((....))))))).(((((((((.(((((((.......)))))))))))).......(((..(.....((.((((((((.....)))))))))))..)))....)))).... ( -30.00) >DroSec_CAF1 11315 120 - 1 AUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUGUCACGAGUUCGUACCCAAAUAUAAAGCGUGGUAUUGGGCUUUAGAAAUAUAUUUUCUGGAUCCUGUUUGAUUGCAAUAUG ..(((((..((..(((((...(((..(((((.(((((((.......)))))))))))).....))).)))))...(((.((((((((.....)))))))).))).....))..))))).. ( -35.50) >DroSim_CAF1 12279 120 - 1 AUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGAAUAUCACGAGUUCGUACCCAAAUAUAAAGCGUGGUAUUGGGCUUUAGAAAUAUAUUUUCUGGAUCCUGUUUGAUUGCAAUAUG ..(((((..((..(((((...(((..(((((.(((((((.......)))))))))))).....))).)))))...(((.((((((((.....)))))))).))).....))..))))).. ( -35.70) >DroEre_CAF1 11046 120 - 1 GUUGUUGUGGGAAACCAUAAUCGUAGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUACAGUGUGCUAUUAGACUUCAGAAUUAUAUGUUCGGGAUCCUGCAUGAUUGUUAUAUG ...(((((((....))))))).....(((((.(((((((.......))))))))))))(((((((((((((.....((.(((.((((.....)))).)))))..)))).)))).))))). ( -33.80) >DroYak_CAF1 13918 120 - 1 GUUGUUGUGGGAAACCAUUACCGUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUAUAGUGUCGUAUUCGUCUUGAGAAAUAUGUGUUCAGGAUCCUGCAAGAUUGCGAUAUG ...((.((((....)))).)).(((.(((((.(((((((.......)))))))))))).)))...((((((((.((((((((((.........)))))))).......)).)))))))). ( -36.01) >consensus GUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUAAAGUGUGGUAUUGGACUUUAGAAAUAUAUUUUCUGGAUCCUGCAUGAUUGCAAUAUG ...(((((((....))))))).(((.(((((.(((((((.......))))))))))))...)))...............((((((((.....)))))))).................... (-25.62 = -25.62 + -0.00)

| Location | 7,914,450 – 7,914,570 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 89.08 |
| Mean single sequence MFE | -39.82 |
| Consensus MFE | -32.84 |
| Energy contribution | -32.72 |
| Covariance contribution | -0.12 |
| Combinations/Pair | 1.12 |
| Mean z-score | -1.34 |
| Structure conservation index | 0.82 |
| SVM decision value | -0.04 |
| SVM RNA-class probability | 0.514479 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
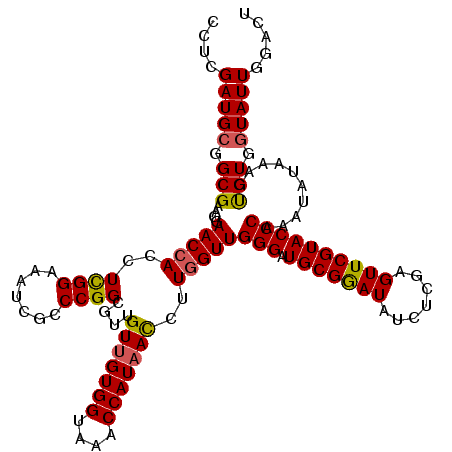
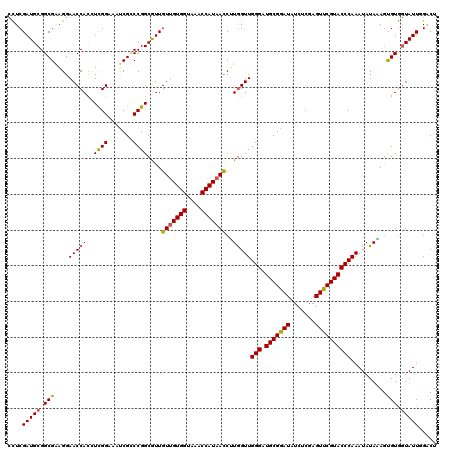
>2R_DroMel_CAF1 7914450 120 - 20766785 CCUCGAUGCGGCGGAGGAACCACAUCGGAAAUCGCCCGGCGUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUACAAUGUGGUAUUUGACU ((((.........)))).((((((((((.......))).((..(((((((....)))))))..)).(((((.(((((((.......)))))))))))).......)))))))........ ( -37.50) >DroSec_CAF1 11355 120 - 1 CCUCGAUGCGGCGAAGGAACCACCUCGGAAAUCGCCCGGCAUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUGUCACGAGUUCGUACCCAAAUAUAAAGCGUGGUAUUGGGCU .((((((((.(((....(((((..((((.......))))....(((((((....)))))))..)))))(((.(((((((.......))))))))))..........))).)))))))).. ( -41.50) >DroSim_CAF1 12319 120 - 1 CCUCGAUGCGGCGAAGGAACCACCUCGGAAAUCGCCCGGCAUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGAAUAUCACGAGUUCGUACCCAAAUAUAAAGCGUGGUAUUGGGCU .((((((((.(((....(((((..((((.......))))....(((((((....)))))))..)))))(((.(((((((.......))))))))))..........))).)))))))).. ( -41.70) >DroEre_CAF1 11086 120 - 1 ACUCGAUGCGGCGAAGGAACCAUUUUGGAAAUCGUCCGGCGUUGUUGUGGGAAACCAUAAUCGUAGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUACAGUGUGCUAUUAGACU ...(((((((((((.....((.....))...))).)).))))))((((((....))))))..(((.(((((.(((((((.......))))))))))))...)))(((.((....)).))) ( -36.60) >DroYak_CAF1 13958 120 - 1 CUUCGAUGCGGCGCAGGAACCAUUUCGGAAACCGGCCGGCGUUGUUGUGGGAAACCAUUACCGUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUAUAGUGUCGUAUUCGUCU ...(((((((((((...((((((.((((.......))))....((.((((....)))).)).))))))(((.(((((((.......)))))))))).........)))))))).)))... ( -41.80) >consensus CCUCGAUGCGGCGAAGGAACCACCUCGGAAAUCGCCCGGCGUUGUUGUGGUAAACCAUAACCUUGGUUGGGAUGCGGAUAUCUCGAGUUCGUACCCAAAUAUAAAGUGUGGUAUUGGACU ....(((((.(((....(((((..((((.......))))....(((((((....)))))))..)))))(((.(((((((.......))))))))))..........))).)))))..... (-32.84 = -32.72 + -0.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:36 2006