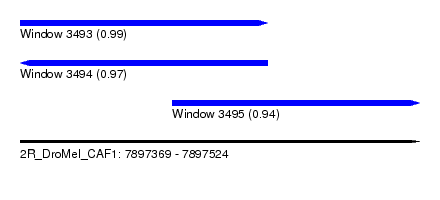

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,897,369 – 7,897,524 |
| Length | 155 |
| Max. P | 0.988971 |

| Location | 7,897,369 – 7,897,465 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | forward |
| Mean pairwise identity | 93.88 |
| Mean single sequence MFE | -29.52 |
| Consensus MFE | -25.06 |
| Energy contribution | -25.38 |
| Covariance contribution | 0.32 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.10 |
| Structure conservation index | 0.85 |
| SVM decision value | 2.14 |
| SVM RNA-class probability | 0.988971 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 7897369 96 + 20766785 GUGACUAAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUGAAGAAACCCUGGUCACUCACUUAAUUGAUUCCAGGCAAUCGAUGGAUCAUCGACGGGG- ((((((...((((((.(((...))).))))))(((((......)))))))))))...(((..(((((((((.........))))..)))))..)))- ( -28.10) >DroSec_CAF1 1819 96 + 1 GUGACUCAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUUAAGAAACCCUGGUCACUCACUUAAUCGAUUCCAGACAAUCGAUGGAUCAUCGAAGUGG- ((((((...((((((.(((...))).))))))((((((....)))))))))))).((((...(((((((((.........))))..))))))))).- ( -31.40) >DroSim_CAF1 1817 96 + 1 GUGACUCAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUUAAGAAACCCUGGUCACUCACUUAAUCGAUUCCAGACAAUCGAUGGAUCAUCGAAGUGG- ((((((...((((((.(((...))).))))))((((((....)))))))))))).((((...(((((((((.........))))..))))))))).- ( -31.40) >DroEre_CAF1 1810 97 + 1 GUGACUAAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUUAAGAAACCCCGGGCCCUCAAUUAAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGGU .(((((...((((((.(((...))).))))))..)))))....((((((((...((((....))))..))......((((((...)))))))))))) ( -30.00) >DroYak_CAF1 1828 96 + 1 GUGACUAAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUUAAGAAACCCCGGCCACUCAAUUAAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGG- .(((((...((((((.(((...))).))))))..))))).....(((((.....((((....))))..........((((((...)))))))))))- ( -26.70) >consensus GUGACUAAAGUGAAUGAUGCUCCAUGAUUCAUGGGGUUAAGAAACCCUGGUCACUCACUUAAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGG_ ((((((...((((((.(((...))).))))))((((((....))))))))))))........(((((((((.........))))..)))))...... (-25.06 = -25.38 + 0.32)

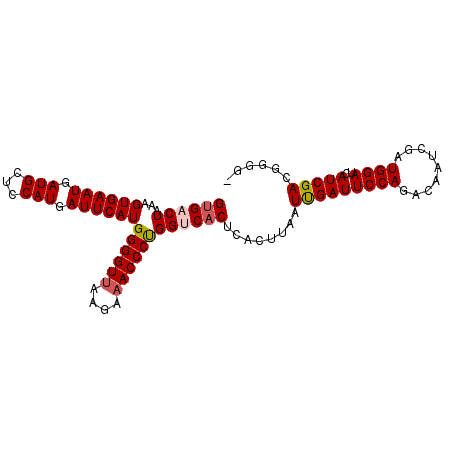
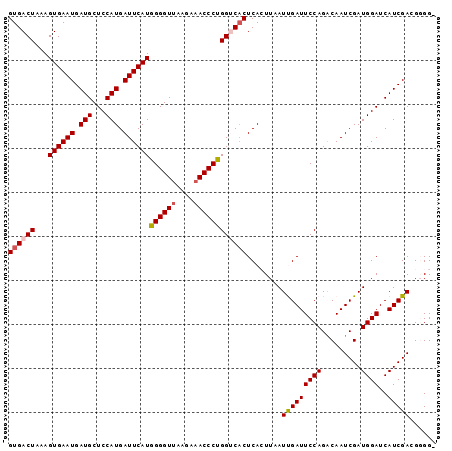
| Location | 7,897,369 – 7,897,465 |
|---|---|
| Length | 96 |
| Sequences | 5 |
| Columns | 97 |
| Reading direction | reverse |
| Mean pairwise identity | 93.88 |
| Mean single sequence MFE | -27.78 |
| Consensus MFE | -22.12 |
| Energy contribution | -21.84 |
| Covariance contribution | -0.28 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.35 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965117 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 7897369 96 - 20766785 -CCCCGUCGAUGAUCCAUCGAUUGCCUGGAAUCAAUUAAGUGAGUGACCAGGGUUUCUUCACCCCAUGAAUCAUGGAGCAUCAUUCACUUUAGUCAC -..(((((((((...))))))).....))........((((((((((...((((......))))((((...)))).....))))))))))....... ( -23.70) >DroSec_CAF1 1819 96 - 1 -CCACUUCGAUGAUCCAUCGAUUGUCUGGAAUCGAUUAAGUGAGUGACCAGGGUUUCUUAACCCCAUGAAUCAUGGAGCAUCAUUCACUUGAGUCAC -(((..((((((...)))))).....)))....((((((((((((((...(((((....)))))((((...)))).....)))))))))))).)).. ( -29.80) >DroSim_CAF1 1817 96 - 1 -CCACUUCGAUGAUCCAUCGAUUGUCUGGAAUCGAUUAAGUGAGUGACCAGGGUUUCUUAACCCCAUGAAUCAUGGAGCAUCAUUCACUUGAGUCAC -(((..((((((...)))))).....)))....((((((((((((((...(((((....)))))((((...)))).....)))))))))))).)).. ( -29.80) >DroEre_CAF1 1810 97 - 1 ACCCCGUCGAUGAUCCAUCGAUUGUCUGGAAUCAAUUAAUUGAGGGCCCGGGGUUUCUUAACCCCAUGAAUCAUGGAGCAUCAUUCACUUUAGUCAC ........((((.(((((.((((....((..((((....))))...)).((((((....))))))...))))))))).))))............... ( -27.70) >DroYak_CAF1 1828 96 - 1 -CCCCGUCGAUGAUCCAUCGAUUGUCUGGAAUCAAUUAAUUGAGUGGCCGGGGUUUCUUAACCCCAUGAAUCAUGGAGCAUCAUUCACUUUAGUCAC -.......((((.(((((.((((....((..((((....))))....))((((((....))))))...))))))))).))))............... ( -27.90) >consensus _CCCCGUCGAUGAUCCAUCGAUUGUCUGGAAUCAAUUAAGUGAGUGACCAGGGUUUCUUAACCCCAUGAAUCAUGGAGCAUCAUUCACUUUAGUCAC ........((((.(((((.((((...(((..(((......)))....)))(((((....)))))....))))))))).))))............... (-22.12 = -21.84 + -0.28)

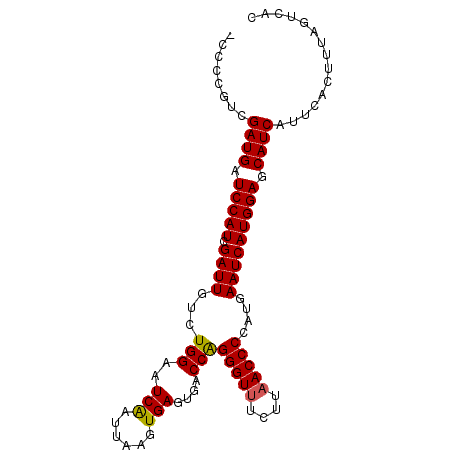
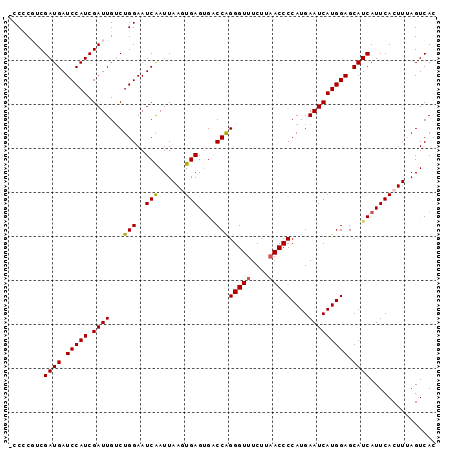
| Location | 7,897,428 – 7,897,524 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 81.81 |
| Mean single sequence MFE | -26.98 |
| Consensus MFE | -15.52 |
| Energy contribution | -16.93 |
| Covariance contribution | 1.42 |
| Combinations/Pair | 1.17 |
| Mean z-score | -3.52 |
| Structure conservation index | 0.57 |
| SVM decision value | 1.25 |
| SVM RNA-class probability | 0.936537 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 7897428 96 + 20766785 U--AAUUGAUUCCAGGCAAUCGAUGGAUCAUCGACGGGG--AAUUAAAAAUUCUAUUAAAAUUGCAACUUGUUGAUAAGCCAACAGGUUAGCAAUGCCCC- .--..((((((((......((((((...))))))...))--)))))).............((((((((((((((......))))))))).))))).....- ( -28.20) >DroPse_CAF1 1859 87 + 1 UAUAAGUGAUUCCGUACGAUCGAUGGAUCGCUUGCAGGAAUGUGUAUGUAUUCUAUUAAAACUGCAA-------------CAACAGGUUAGCAUUUAUCU- .(((((((...((...(((((....))))).(((((((((((......)))))........))))))-------------.....))....)))))))..- ( -18.50) >DroSec_CAF1 1878 96 + 1 U--AAUCGAUUCCAGACAAUCGAUGGAUCAUCGAAGUGG--AAUUAAAAAUUCUAUUAAAAUUGCAACUUGUUGAUAAGCCAACAGGUUAGCAAUGCUCC- .--..(((((((((.........))))..)))))(((((--(((.....))))))))...((((((((((((((......))))))))).))))).....- ( -27.80) >DroSim_CAF1 1876 96 + 1 U--AAUCGAUUCCAGACAAUCGAUGGAUCAUCGAAGUGG--AAUUAAAAAUUCUAUUAAAAUUGCAACUUGUUGAUAAGCCAACAGGUUAGCAAUGCUCC- .--..(((((((((.........))))..)))))(((((--(((.....))))))))...((((((((((((((......))))))))).))))).....- ( -27.80) >DroEre_CAF1 1869 98 + 1 U--AAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGGUCGAUUAAGAAUUCUAUUAAAACUGCAACUUGUUGAUAAGUCAACAGGUUAGCAAUUCUCC- (--((((((((((......((((((...)))))).)))))))))))................(((((((((((((....)))))))))).))).......- ( -29.40) >DroYak_CAF1 1887 97 + 1 U--AAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGG--AAUUAAGAAUUCUAUUAAAAUUGCAACUUGUUGAUAAGCCAACAGGUUAGCAAUUUUCAC .--..((((((((......((((((...))))))...))--))))))..........(((((((((((((((((......))))))))).))))))))... ( -30.20) >consensus U__AAUUGAUUCCAGACAAUCGAUGGAUCAUCGACGGGG__AAUUAAAAAUUCUAUUAAAAUUGCAACUUGUUGAUAAGCCAACAGGUUAGCAAUGCUCC_ .....(((((((((.........))))..)))))..........................((((((((((((((......))))))))).)))))...... (-15.52 = -16.93 + 1.42)

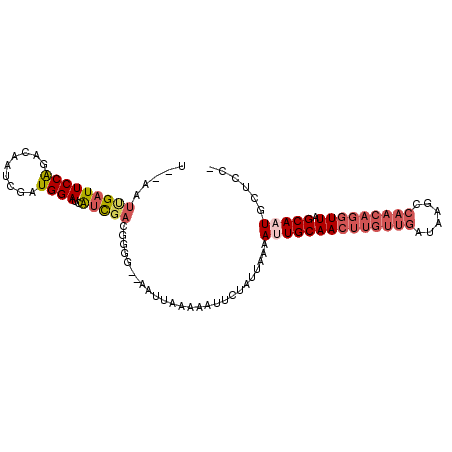
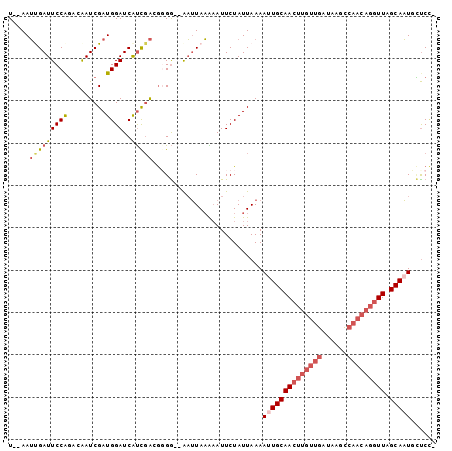
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:21:30 2006