
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,517,725 – 7,517,820 |
| Length | 95 |
| Max. P | 0.884782 |

| Location | 7,517,725 – 7,517,820 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -21.42 |
| Consensus MFE | -15.37 |
| Energy contribution | -15.23 |
| Covariance contribution | -0.14 |
| Combinations/Pair | 1.18 |
| Mean z-score | -1.80 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.90 |
| SVM RNA-class probability | 0.878519 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
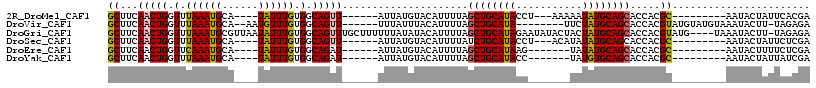
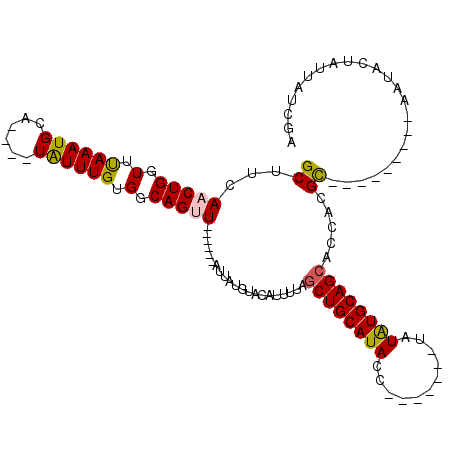
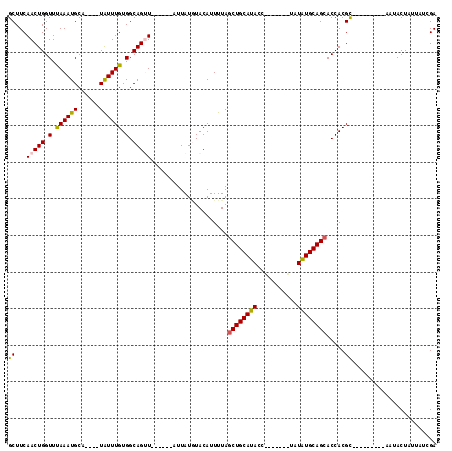
>2R_DroMel_CAF1 7517725 95 + 20766785 GCUUCAACUGGUUUAAAUGCA----UAUUUGUGGCAGUU------AUUAUGUACAUUUUAGCUGCAUACCU---AAAAAUAUGCAGCACCACGC---------AAUACUAUUCACGA ........((((.....((((----((((((((((((((------(..((....))..))))))).)))..---..)))))))))..))))...---------.............. ( -21.90) >DroVir_CAF1 22400 100 + 1 GCUUCAACUGGUUUAAAUGCA--AAUGUUUGUGGCAGUU------UUUAUUUACAUUUUAGCUGCAUA--------UUCUAUGCAGCACCACGUAUGUAUGUAAAUACUU-UAGAGA .......((((..((..((((--.((((.(((((..((.------.......))......((((((((--------...)))))))).))))).)))).))))..))..)-)))... ( -22.80) >DroGri_CAF1 18500 112 + 1 GCUUCAACUGGUUUAAAUGCGUUAAUAUUUGUGGCAGUUUGCUUUUUUAUAUACAUUUUAGCUGCAUAGAAUAUACUACUAUGCAGCACCACGUAUG----UAAAUACUU-UAGAGA ((...(((((.(.((((((......)))))).).))))).))...(((((((((......(((((((((.........))))))))).....)))))----)))).....-...... ( -27.70) >DroSec_CAF1 13066 95 + 1 GCUUCAACUGGUUUAAAUGCA----UAUUUGUGGCAGUU------AUUAUGUACAUUUUAUCUGCAUACCU---ACAUAUAUGCAGCACCACGC---------AAUACUAUUCUCGA ........((((.....((((----(((.((((((((.(------(..((....))..)).)))).....)---))).)))))))..))))...---------.............. ( -17.90) >DroEre_CAF1 13855 91 + 1 GCUUCAACUGGUUCAAAUGCA----UAUUUGUGGCAGAU------AUUAUGUACAUUUUAGCUGCAUAAG-------UAUAUGCAGCACCACGC---------AAUACUUUUCUCGA ((.....(((.(.(((((...----.))))).).)))..------...............((((((((..-------..)))))))).....))---------.............. ( -19.40) >DroYak_CAF1 13359 91 + 1 GCUUCAACUGGUUUAAAUGCA----UAUUUGUGGCAGAU------AUUAUGUACAUUUUAGCUGCAUACC-------UAUGUGCAGCACCACGC---------AAUACUAUUAUCGA ........((((.....((((----(((..(((((((.(------(..((....))..)).)))).))).-------.)))))))..))))...---------.............. ( -18.80) >consensus GCUUCAACUGGUUUAAAUGCA____UAUUUGUGGCAGUU______AUUAUGUACAUUUUAGCUGCAUACC_______UAUAUGCAGCACCACGC_________AAUACUAUUAUCGA ((...(((((.(.((((((......)))))).).))))).....................((((((((...........)))))))).....))....................... (-15.37 = -15.23 + -0.14)
| Location | 7,517,725 – 7,517,820 |
|---|---|
| Length | 95 |
| Sequences | 6 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 79.57 |
| Mean single sequence MFE | -21.25 |
| Consensus MFE | -15.65 |
| Energy contribution | -16.02 |
| Covariance contribution | 0.36 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.74 |
| SVM decision value | 0.93 |
| SVM RNA-class probability | 0.884782 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
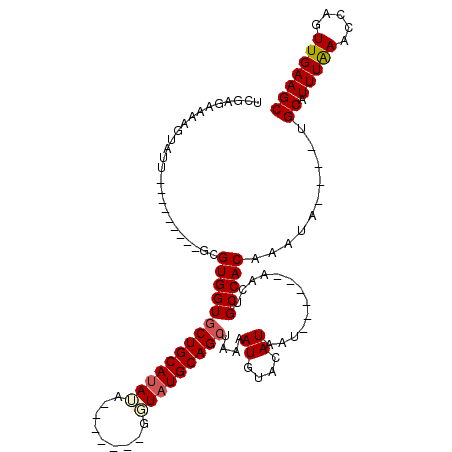
>2R_DroMel_CAF1 7517725 95 - 20766785 UCGUGAAUAGUAUU---------GCGUGGUGCUGCAUAUUUUU---AGGUAUGCAGCUAAAAUGUACAUAAU------AACUGCCACAAAUA----UGCAUUUAAACCAGUUGAAGC ...(((((.(((((---------..(((((((((((((((...---.))))))))))......((.......------.)).))))).))))----)..)))))............. ( -23.80) >DroVir_CAF1 22400 100 - 1 UCUCUA-AAGUAUUUACAUACAUACGUGGUGCUGCAUAGAA--------UAUGCAGCUAAAAUGUAAAUAAA------AACUGCCACAAACAUU--UGCAUUUAAACCAGUUGAAGC ......-..((((....)))).....((((((((((((...--------))))))))..((((((((((...------.............)))--)))))))..))))........ ( -20.69) >DroGri_CAF1 18500 112 - 1 UCUCUA-AAGUAUUUA----CAUACGUGGUGCUGCAUAGUAGUAUAUUCUAUGCAGCUAAAAUGUAUAUAAAAAAGCAAACUGCCACAAAUAUUAACGCAUUUAAACCAGUUGAAGC ......-.....((((----.((((((...(((((((((.........)))))))))....)))))).))))...((.(((((....((((........))))....)))))...)) ( -23.70) >DroSec_CAF1 13066 95 - 1 UCGAGAAUAGUAUU---------GCGUGGUGCUGCAUAUAUGU---AGGUAUGCAGAUAAAAUGUACAUAAU------AACUGCCACAAAUA----UGCAUUUAAACCAGUUGAAGC ....((((.(((((---------..(((((.((((((((....---..)))))))).......((.......------.)).))))).))))----)..)))).............. ( -19.20) >DroEre_CAF1 13855 91 - 1 UCGAGAAAAGUAUU---------GCGUGGUGCUGCAUAUA-------CUUAUGCAGCUAAAAUGUACAUAAU------AUCUGCCACAAAUA----UGCAUUUGAACCAGUUGAAGC (((((....(((((---------..(((((((((((((..-------..))))))))....((((.....))------))..))))).))))----)...)))))............ ( -22.70) >DroYak_CAF1 13359 91 - 1 UCGAUAAUAGUAUU---------GCGUGGUGCUGCACAUA-------GGUAUGCAGCUAAAAUGUACAUAAU------AUCUGCCACAAAUA----UGCAUUUAAACCAGUUGAAGC ((((((((.(((((---------..(((((((((((....-------....))))))....((((.....))------))..))))).))))----)..))).......)))))... ( -17.41) >consensus UCGAGAAAAGUAUU_________GCGUGGUGCUGCAUAUA_______GGUAUGCAGCUAAAAUGUACAUAAU______AACUGCCACAAAUA____UGCAUUUAAACCAGUUGAAGC .........................((((((((((((((.........)))))))))....((....)).............)))))..........((.(((((.....))))))) (-15.65 = -16.02 + 0.36)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:18:15 2006