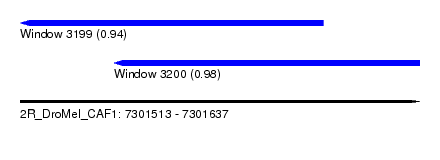

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 7,301,513 – 7,301,637 |
| Length | 124 |
| Max. P | 0.978710 |

| Location | 7,301,513 – 7,301,607 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 88.71 |
| Mean single sequence MFE | -29.13 |
| Consensus MFE | -22.90 |
| Energy contribution | -23.14 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.30 |
| SVM RNA-class probability | 0.939805 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 7301513 94 - 20766785 CCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCCAGCCAACGGCGUGGGUGGUCAUUUUAACCUCUUGGCCAACUACUC- (((((..(((((((((....))))).-.................((((....)))))))))))))((((((...........)))))).......- ( -27.60) >DroSim_CAF1 101221 94 - 1 CCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCUAGCCGACGGCGUGGGUGGUCAUUUUAACCUCUUGGCCAACUACAC- (((((..((((((...((((((..(.-.................)..))).))))))))))))))((((((...........)))))).......- ( -30.07) >DroEre_CAF1 97432 94 - 1 CCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGUGGUCAUUUUAGCCUCUUGGCCAACUACUC- (((((..(((((((((....)))...-...(((..((((.....))))..))).)))))))))))((((((...........)))))).......- ( -28.50) >DroYak_CAF1 99710 94 - 1 CCCAUUUGCCGUCAACGGCAGUGGAA-AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGUGGCCAUUUUAACCUCUUGGCCAACUACUC- (((((..((((((...(((((((...-..))))..((((.....))))...))))))))))))))((((((...........)))))).......- ( -33.30) >DroAna_CAF1 91690 81 - 1 CCCAUUUGCCGUCAACGGCAGCCGAAAAACGCUCAUCAAACUAAUUGGCCAG-------CGUGGGUGGUC--------UUCUUGGCCAACUACAUG (((..((((((....)))))).......(((((..((((.....))))..))-------))))))(((((--------.....)))))........ ( -26.20) >consensus CCCAUUUGCCGUCAACGGCAGUUGAA_AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGUGGUCAUUUUAACCUCUUGGCCAACUACUC_ (((((((((((....)))))).........(((..((((.....))))..))).......)))))((((((...........))))))........ (-22.90 = -23.14 + 0.24)


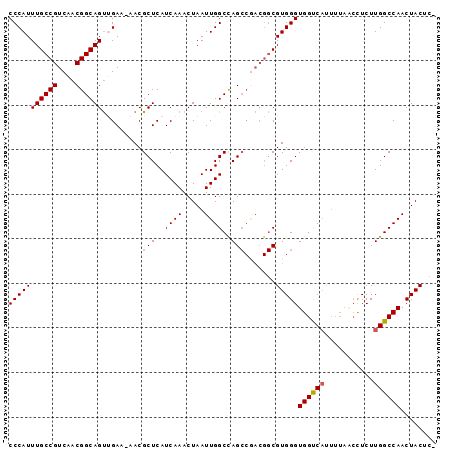
| Location | 7,301,542 – 7,301,637 |
|---|---|
| Length | 95 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 91.58 |
| Mean single sequence MFE | -28.65 |
| Consensus MFE | -22.19 |
| Energy contribution | -22.13 |
| Covariance contribution | -0.06 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.77 |
| SVM decision value | 1.82 |
| SVM RNA-class probability | 0.978710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 7301542 95 - 20766785 CCGCAAAAAGCUAAUUAAUUUUUU-----GCUCUACCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCCAGCCAACGGCGUGGGU ..((((((((........))))))-----))...((((((..(((((((((....))))).-.................((((....)))))))))))))) ( -28.10) >DroSim_CAF1 101250 95 - 1 CCGCAAAAAGCUAAUUAAUUUUUU-----GCUCUACCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCUAGCCGACGGCGUGGGU ..((((((((........))))))-----))...((((((..((((((...((((((..(.-.................)..))).))))))))))))))) ( -30.57) >DroEre_CAF1 97461 95 - 1 CCGCAAAAAGCUAAUUAAUUUUUU-----GCUCUACCCAUUUGCCGUCAACGGCAGUUGAA-AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGU ..((((((((........))))))-----))...((((((..(((((((((....)))...-...(((..((((.....))))..))).)))))))))))) ( -29.00) >DroYak_CAF1 99739 95 - 1 CCGCAAAAAGCUAAUUAAUUUUUU-----GCUCUACCCAUUUGCCGUCAACGGCAGUGGAA-AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGU ..((((((((........))))))-----))...((((((..((((((...(((((((...-..))))..((((.....))))...))))))))))))))) ( -31.10) >DroAna_CAF1 91712 94 - 1 CCGCAAAAAGUUAAUUAAUUUUUUCCCUUGCACUGCCCAUUUGCCGUCAACGGCAGCCGAAAAACGCUCAUCAAACUAAUUGGCCAG-------CGUGGGU ..((((((((((....)))))).....))))...((((..((((((....)))))).......(((((..((((.....))))..))-------))))))) ( -24.50) >consensus CCGCAAAAAGCUAAUUAAUUUUUU_____GCUCUACCCAUUUGCCGUCAACGGCAGUUGAA_AACGCUCAUCAAACUAAUUGGCCAGCCGACGGCGUGGGU ((((.....((((((((.(((........((.........((((((....)))))).........)).....))).))))))))..(((...))))))).. (-22.19 = -22.13 + -0.06)


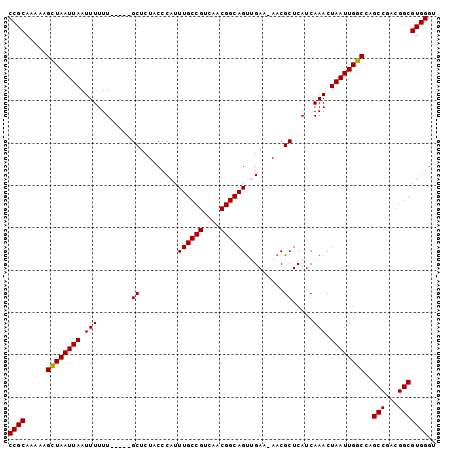
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:16:45 2006