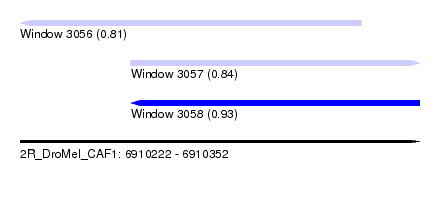

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,910,222 – 6,910,352 |
| Length | 130 |
| Max. P | 0.929510 |

| Location | 6,910,222 – 6,910,333 |
|---|---|
| Length | 111 |
| Sequences | 6 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 81.18 |
| Mean single sequence MFE | -34.68 |
| Consensus MFE | -27.85 |
| Energy contribution | -27.85 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.80 |
| SVM decision value | 0.66 |
| SVM RNA-class probability | 0.814928 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
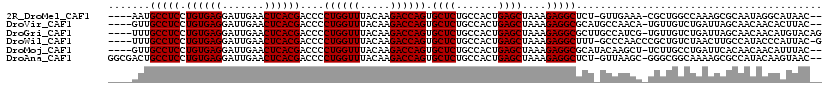
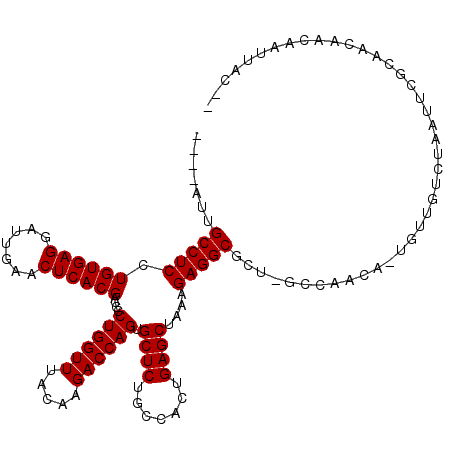
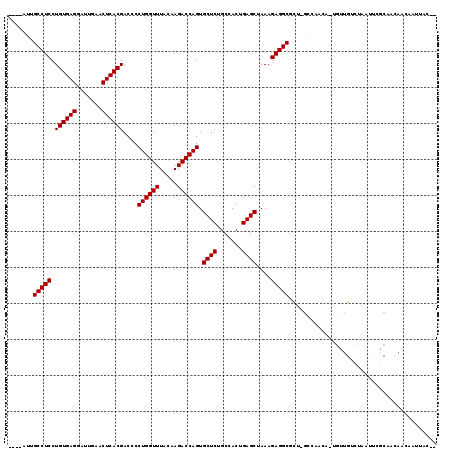
>2R_DroMel_CAF1 6910222 111 - 20766785 ----AAUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCUCU-GUUGAAA-CGCUGGCCAAAGCGCAAUAGGCAUAAC-- ----...(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))(((-((((...-((((......)))))))))))......-- ( -37.40) >DroVir_CAF1 34435 112 - 1 ----GUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCGCAUGCCAACA-UGUUGUCUGAUUAGCAACAACACUUAC-- ----((.(((((.((((((.......))))))....((((((.....)))))).((((.......))))....))))))).........-((((((.......))))))........-- ( -32.90) >DroGri_CAF1 34833 114 - 1 ----UUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCGCUUGCCAUCG-UGUUGUCUGAUUAGCAACAACAUGUACAG ----...(((((.((((((.......))))))....((((((.....)))))).((((.......))))....))))).........((-((((((.((.....))))))))))..... ( -36.00) >DroWil_CAF1 41712 113 - 1 ----UUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCUUU-GCCCAACCCGCUGUCUAACUUGCCAUACCCAUUAC-G ----...(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))...-...................................-. ( -27.60) >DroMoj_CAF1 39343 112 - 1 ----GUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCGCAUACAAGCU-UCUUGCCUGAUUCACAACAACAUUUAC-- ----((((((((.((((((.......))))))....((((((.....)))))).((((.......))))....))))).....((.((.-....)).))......))).........-- ( -31.20) >DroAna_CAF1 30191 115 - 1 GGCGACUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCUCU-GUUAAGC-GGGCGGCAAAAGCGCCAUACAAGUAAC-- ((((..((((....(((((.......)))))(.(((((((((.....)))))).(((..((....(((((.....))))).-))..)))-))))))))....))))...........-- ( -43.00) >consensus ____AUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCGCU_GCCAACA_UGUUGUCUAAUUCGCAACAACAAUUAC__ .......(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))......................................... (-27.85 = -27.85 + -0.00)
| Location | 6,910,258 – 6,910,352 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | forward |
| Mean pairwise identity | 80.44 |
| Mean single sequence MFE | -29.47 |
| Consensus MFE | -27.60 |
| Energy contribution | -27.60 |
| Covariance contribution | -0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.52 |
| Structure conservation index | 0.94 |
| SVM decision value | 0.74 |
| SVM RNA-class probability | 0.838568 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
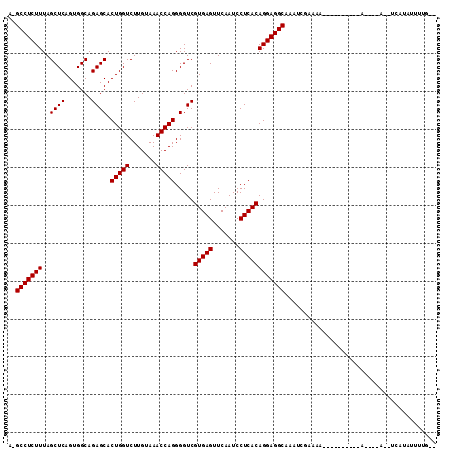
>2R_DroMel_CAF1 6910258 94 + 20766785 A-GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAUUUGUACGA----------A----A--UCAUGUUUUU-- .-(((((((..((((.......)))).(((((.......))))).....(((((.......))))))))))))..........----------.----.--..........-- ( -27.80) >DroVir_CAF1 34472 103 + 1 C-GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAACUCGAAAA-ACUCGUGCAA----A--UCGUUUUUUU-- .-(((((((..((((.......)))).(((((.......))))).....(((((.......)))))))))))).....(((((-((........----.--..))))))).-- ( -29.70) >DroGri_CAF1 34872 97 + 1 C-GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAAACUAAAAAGAU-C----------A--AUACAUUUUG-- .-(((((((..((((.......)))).(((((.......))))).....(((((.......)))))))))))).............-.----------.--..........-- ( -27.90) >DroWil_CAF1 41750 102 + 1 A-GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAAAAAUAUCC----------AUCAAAGUUCAAACUUUGUG .-(((((((..((((.......)))).(((((.......))))).....(((((.......)))))))))))).........(----------(.((((((....)))))))) ( -31.50) >DroMoj_CAF1 39380 104 + 1 C-GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAACUCGAAAAAACGUUUGGGA----A--AUAUUUUUUG-- .-(((((((..((((.......)))).(((((.......))))).....(((((.......))))))))))))..((((((......)))))).----.--..........-- ( -31.60) >DroPer_CAF1 27465 92 + 1 AGGCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAAUAGC---C----------A----A--UUUUCUUUUU-- .(((((.....((((.......)))).(((((.......))))))))))(((((.......)))))....(((....))---)----------.----.--..........-- ( -28.30) >consensus A_GCCUCUUUAGCUCAGUGGCAGAGCACUGGUCUUGUAAACCAGGGGUCGUGAGUUCAAUCCUCACAGGAGGCAAAUCGAAAA__________A____A__UCAUAUUUUG__ ..(((((((..((((.......)))).(((((.......))))).....(((((.......))))))))))))........................................ (-27.60 = -27.60 + -0.00)


| Location | 6,910,258 – 6,910,352 |
|---|---|
| Length | 94 |
| Sequences | 6 |
| Columns | 113 |
| Reading direction | reverse |
| Mean pairwise identity | 80.44 |
| Mean single sequence MFE | -29.05 |
| Consensus MFE | -26.53 |
| Energy contribution | -26.53 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.92 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.20 |
| SVM RNA-class probability | 0.929510 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6910258 94 - 20766785 --AAAAACAUGA--U----U----------UCGUACAAAUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC-U --..........--.----.----------..........(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))-. ( -27.60) >DroVir_CAF1 34472 103 - 1 --AAAAAAACGA--U----UUGCACGAGU-UUUUCGAGUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC-G --.......(((--.----..((....))-...)))....(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))-. ( -30.90) >DroGri_CAF1 34872 97 - 1 --CAAAAUGUAU--U----------G-AUCUUUUUAGUUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC-G --..........--.----------.-.............(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))-. ( -28.10) >DroWil_CAF1 41750 102 - 1 CACAAAGUUUGAACUUUGAU----------GGAUAUUUUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC-U ((((((((....)))))).)----------).........(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))-. ( -32.30) >DroMoj_CAF1 39380 104 - 1 --CAAAAAAUAU--U----UCCCAAACGUUUUUUCGAGUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC-G --..........--.----.......((......))....(((((.((((((.......))))))....((((((.....)))))).((((.......))))....)))))-. ( -28.20) >DroPer_CAF1 27465 92 - 1 --AAAAAGAAAA--U----U----------G---GCUAUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGCCU --..........--.----.----------.---......(((((.((((((.......))))))....((((((.....)))))).((((.......))))....))))).. ( -27.20) >consensus __AAAAAAAUAA__U____U__________UUUUACAAUUGCCUCCUGUGAGGAUUGAACUCACGACCCCUGGUUUACAAGACCAGUGCUCUGCCACUGAGCUAAAGAGGC_G ........................................(((((.((((((.......))))))....((((((.....)))))).((((.......))))....))))).. (-26.53 = -26.53 + 0.00)

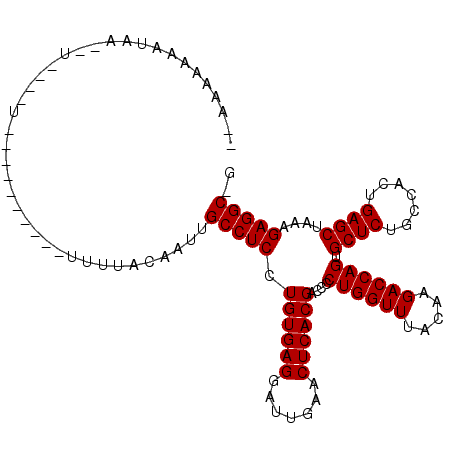
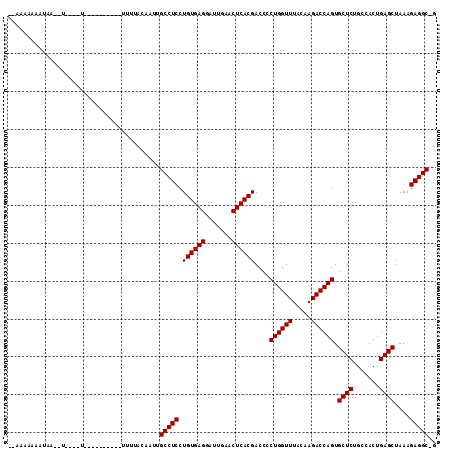
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:14:31 2006