
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,892,569 – 6,892,683 |
| Length | 114 |
| Max. P | 0.994850 |

| Location | 6,892,569 – 6,892,683 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 93.27 |
| Mean single sequence MFE | -58.33 |
| Consensus MFE | -53.71 |
| Energy contribution | -53.90 |
| Covariance contribution | 0.19 |
| Combinations/Pair | 1.09 |
| Mean z-score | -3.15 |
| Structure conservation index | 0.92 |
| SVM decision value | 2.52 |
| SVM RNA-class probability | 0.994850 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6892569 114 + 20766785 AAGCAGCGGGAGCAGGAGGAGGCCCGAGUUCUCGAGGAGGCGGAGGCCGCCGAAGCGGCUGCUGCAGCCGCAAUCCUUGAGAGCGGCGAAAAGGAUGGUCGCAAGCCUCCAGUC .....((....))....((((((....((((((((((((((((...)))))...(((((((...)))))))..))))))))))).((((.........))))..)))))).... ( -59.90) >DroSec_CAF1 8336 114 + 1 AAGCAGCGGGAGCAGGAGGAGGCCCGGAUCCUCGAGGAGGCGGAGGCCGCCGAAGCGGCUGCUGCAGCCGCAAUCCUCGAGAGCGGCGAAAAGGAGGGUCGCAAGCCUCCAGUC .....((....))....((((((.......(((((((((((((...)))))...(((((((...)))))))..)))))))).(((((..........)))))..)))))).... ( -59.00) >DroSim_CAF1 12482 114 + 1 AAGCAGCGGGAGCAGGAGGAGGCCCGGGUACUCGAGGAGGCGGAGGCUGCCGAAGCGGCUGCUGCAGCCGCAAUCCUCGAGAGCGGCGAAAAGGAGGGUCGCAAGCCACCAUUC .....((....))....((.(((.......(((((((((((((...)))))...(((((((...)))))))..)))))))).(((((..........)))))..))).)).... ( -52.10) >DroEre_CAF1 12944 114 + 1 AAGCAGCGCGAGCAGGAGGAGGCCCGGGUUCUCGAGGAGGCGGAGGCCGCUGAGGCGGCCGCUGCAGCCGCCAUCCUCGAGAGCGGCGAGAAGGAGGGUCGCAAGCCCCCAGUC .....((....))........(((...((((((((((((((((.((((((....)))))).......))))).)))))))))))))).....((.(((.(....)))))).... ( -62.30) >consensus AAGCAGCGGGAGCAGGAGGAGGCCCGGGUUCUCGAGGAGGCGGAGGCCGCCGAAGCGGCUGCUGCAGCCGCAAUCCUCGAGAGCGGCGAAAAGGAGGGUCGCAAGCCUCCAGUC .....((....))....((((((.......(((((((((((((...)))))...(((((((...)))))))..)))))))).(((((..........)))))..)))))).... (-53.71 = -53.90 + 0.19)

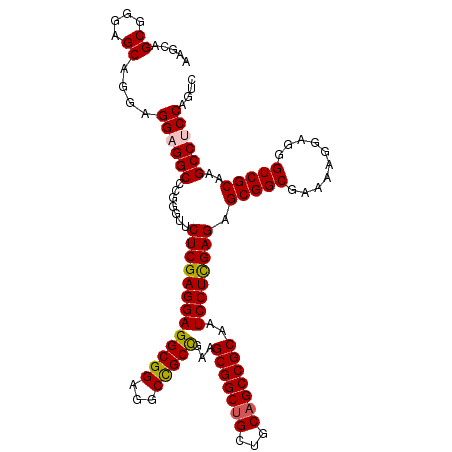
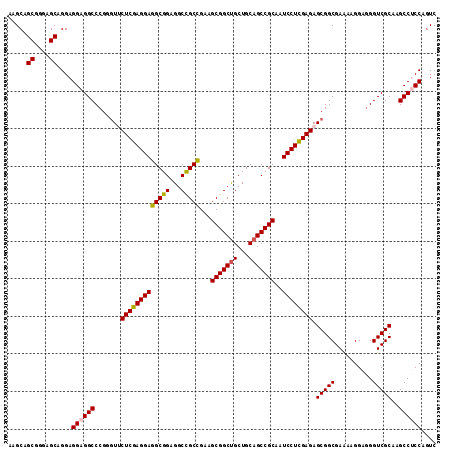
| Location | 6,892,569 – 6,892,683 |
|---|---|
| Length | 114 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 93.27 |
| Mean single sequence MFE | -49.80 |
| Consensus MFE | -46.16 |
| Energy contribution | -46.98 |
| Covariance contribution | 0.81 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.06 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.960387 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6892569 114 - 20766785 GACUGGAGGCUUGCGACCAUCCUUUUCGCCGCUCUCAAGGAUUGCGGCUGCAGCAGCCGCUUCGGCGGCCUCCGCCUCCUCGAGAACUCGGGCCUCCUCCUGCUCCCGCUGCUU ....((((((((((((.........))))...((((.((((..(((((((...)))))))...(((((...))))))))).))))....))))))))....((.......)).. ( -47.70) >DroSec_CAF1 8336 114 - 1 GACUGGAGGCUUGCGACCCUCCUUUUCGCCGCUCUCGAGGAUUGCGGCUGCAGCAGCCGCUUCGGCGGCCUCCGCCUCCUCGAGGAUCCGGGCCUCCUCCUGCUCCCGCUGCUU ....((((((((((((.........))))...(((((((((..(((((((...)))))))...(((((...))))))))))))))....))))))))....((.......)).. ( -52.70) >DroSim_CAF1 12482 114 - 1 GAAUGGUGGCUUGCGACCCUCCUUUUCGCCGCUCUCGAGGAUUGCGGCUGCAGCAGCCGCUUCGGCAGCCUCCGCCUCCUCGAGUACCCGGGCCUCCUCCUGCUCCCGCUGCUU ....((.(((((((((.........)))).(..((((((((..(((((((...)))))))...(((.......)))))))))))..)..))))).))....((.......)).. ( -43.10) >DroEre_CAF1 12944 114 - 1 GACUGGGGGCUUGCGACCCUCCUUCUCGCCGCUCUCGAGGAUGGCGGCUGCAGCGGCCGCCUCAGCGGCCUCCGCCUCCUCGAGAACCCGGGCCUCCUCCUGCUCGCGCUGCUU ((..((((((....).)))))..))..((.(((((((((((.((((((....))((((((....))))))..)))))))))))))...(((((........))))).)).)).. ( -55.70) >consensus GACUGGAGGCUUGCGACCCUCCUUUUCGCCGCUCUCGAGGAUUGCGGCUGCAGCAGCCGCUUCGGCGGCCUCCGCCUCCUCGAGAACCCGGGCCUCCUCCUGCUCCCGCUGCUU ....((((((((((((.........))))...(((((((((..(((((((...)))))))...(((((...))))))))))))))....))))))))....((.......)).. (-46.16 = -46.98 + 0.81)

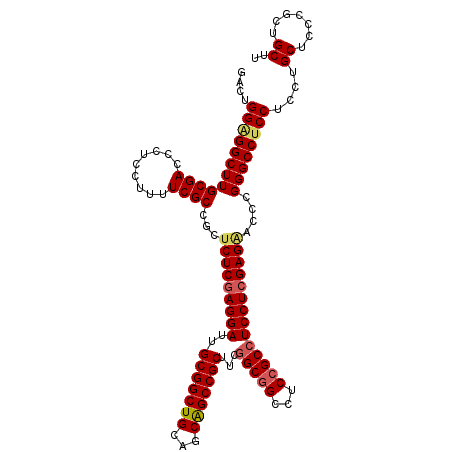
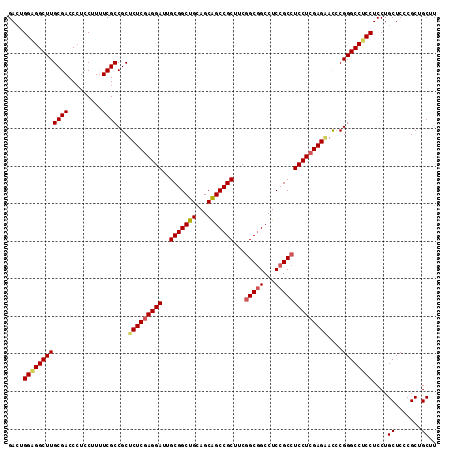
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:14:26 2006