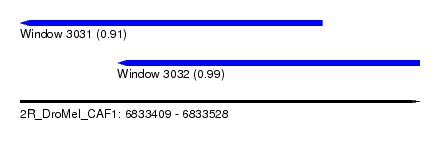

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,833,409 – 6,833,528 |
| Length | 119 |
| Max. P | 0.992459 |

| Location | 6,833,409 – 6,833,499 |
|---|---|
| Length | 90 |
| Sequences | 6 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 80.31 |
| Mean single sequence MFE | -19.68 |
| Consensus MFE | -14.81 |
| Energy contribution | -15.07 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.25 |
| Mean z-score | -2.07 |
| Structure conservation index | 0.75 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.909071 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6833409 90 - 20766785 AUCCAAAAUCCAAAAGCCAGGUUGCGGUGAA-AAUGCCGCGAUCAAGCUAGAAAACCCCUUU--CCAAGUACUCAAAAUCAAA-CCAUUGUGGA ........((((..(((..(((((((((...-...)))))))))..))).((((.....)))--)..................-......)))) ( -20.90) >DroPse_CAF1 7856 78 - 1 ACCCCA-----------CAGAUCGAGGCGAAUAAUGCCGCGAUCCAGCUAGGAAACCCCCACAGAGCAGAGACCGUAAUCGAAACGAUU----- .....(-----------(.(((((.((((.....)))).)))))..(((.((......))....))).......))(((((...)))))----- ( -16.50) >DroSec_CAF1 6317 90 - 1 AUCCAAAAUCCAAAAGCCAGGUUGCGGUGAA-AAUGCCGCGAUCAAGCUAGAAAAUCCCUUU--CCAAGUACUCCAAAUCAAA-CCAUUGUGGA ........((((..(((..(((((((((...-...)))))))))..))).((((.....)))--)..................-......)))) ( -20.90) >DroSim_CAF1 4810 90 - 1 AUCCAAAAUCCAAAAGCCAGGUUGCGGUGAA-AAUGCCGCGAUCAAGCUAGAAAACCCCUUU--CCAAGUACUCCAAAUCAAA-CCAUUGUGGA ........((((..(((..(((((((((...-...)))))))))..))).((((.....)))--)..................-......)))) ( -20.90) >DroEre_CAF1 6975 89 - 1 ACCCA-AAUCCCAAAGCCAGGUUGCGGUGAA-AAUGCCGCGAUCCAGCUAGAAAACCCCUUU--CCAAGUACUCCAAAUCGAA-CCAUUGUGGA .....-..(((((((((..(((((((((...-...)))))))))..))).((((.....)))--)..................-...))).))) ( -21.50) >DroYak_CAF1 7169 84 - 1 ACCCA-AAUCCAAAAGCCAGGUUGCGGUGAA-AAUGCCGCGAUCAAGCUAGAAAACCCCUUU--CCAAGUACUCCAAAUCGAA-CA-U----AA .....-........(((..(((((((((...-...)))))))))..))).((((.....)))--)..................-..-.----.. ( -17.40) >consensus ACCCAAAAUCCAAAAGCCAGGUUGCGGUGAA_AAUGCCGCGAUCAAGCUAGAAAACCCCUUU__CCAAGUACUCCAAAUCAAA_CCAUUGUGGA ..............(((..(((((((((.......)))))))))..)))............................................. (-14.81 = -15.07 + 0.25)

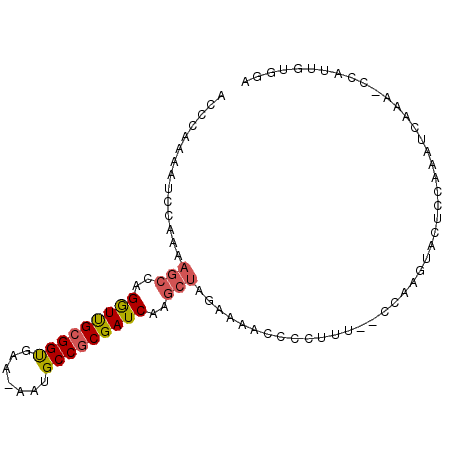

| Location | 6,833,438 – 6,833,528 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 90 |
| Reading direction | reverse |
| Mean pairwise identity | 94.88 |
| Mean single sequence MFE | -19.08 |
| Consensus MFE | -18.04 |
| Energy contribution | -18.24 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.95 |
| SVM decision value | 2.33 |
| SVM RNA-class probability | 0.992459 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6833438 90 - 20766785 CUAAACGCUAAACACGCUAUAGUUUCGAUAUCCAAAAUCCAAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCAAGCUAGAAAACCCCUUU ......((.......)).....(((((((.......)))....(((..(((((((((......)))))))))..))).))))........ ( -18.90) >DroSec_CAF1 6346 90 - 1 CUAAACGCUAAACACGCAAUAGUUUCGAUAUCCAAAAUCCAAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCAAGCUAGAAAAUCCCUUU ......((.......)).....(((((((.......)))....(((..(((((((((......)))))))))..))).))))........ ( -19.40) >DroSim_CAF1 4839 90 - 1 CUAAACGCUAAGCACGCAAUAGUUUCGAUAUCCAAAAUCCAAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCAAGCUAGAAAACCCCUUU ......((.......)).....(((((((.......)))....(((..(((((((((......)))))))))..))).))))........ ( -19.40) >DroEre_CAF1 7004 89 - 1 CUAAACGCCAAACACGCAAUAGUUUCCAUACCCA-AAUCCCAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCCAGCUAGAAAACCCCUUU ......((.......)).................-........(((..(((((((((......)))))))))..)))............. ( -17.80) >DroYak_CAF1 7193 89 - 1 CUAAACGCUAAGCACGCAACAGUUUCGAUACCCA-AAUCCAAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCAAGCUAGAAAACCCCUUU ......((.......)).....(((((((.....-.)))....(((..(((((((((......)))))))))..))).))))........ ( -19.90) >consensus CUAAACGCUAAACACGCAAUAGUUUCGAUAUCCAAAAUCCAAAAGCCAGGUUGCGGUGAAAAUGCCGCGAUCAAGCUAGAAAACCCCUUU ......((.......)).....(((((((.......)))....(((..(((((((((......)))))))))..))).))))........ (-18.04 = -18.24 + 0.20)


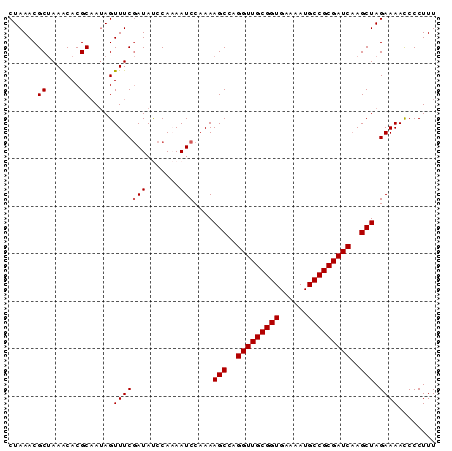
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:14:06 2006