
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,809,527 – 6,809,618 |
| Length | 91 |
| Max. P | 0.968131 |

| Location | 6,809,527 – 6,809,618 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 82.80 |
| Mean single sequence MFE | -17.63 |
| Consensus MFE | -13.60 |
| Energy contribution | -14.93 |
| Covariance contribution | 1.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.75 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.31 |
| SVM RNA-class probability | 0.683347 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
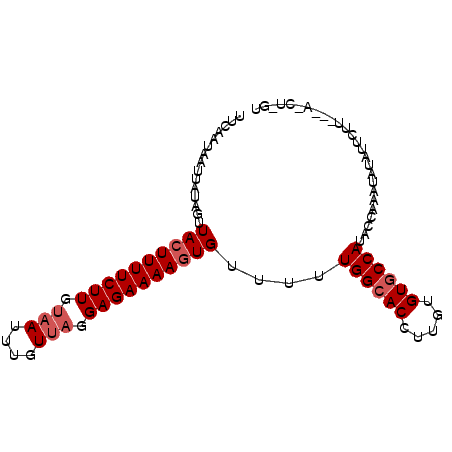
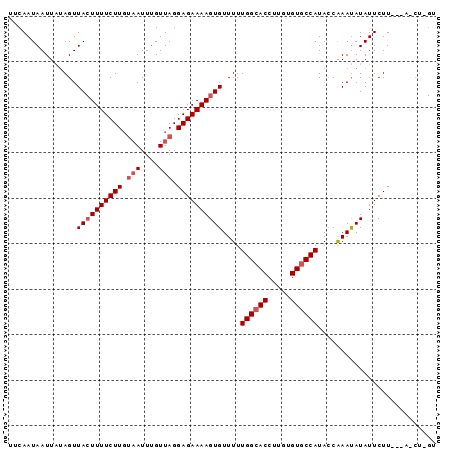
>2R_DroMel_CAF1 6809527 91 + 20766785 AUCAAUAAUUAUAGUUACUUUUCUUGAUAUUUGUUAGGAGAAAAGUGUUUUUGGCACCUUGUGUACCAUAACGAAUAUAUUCUU---AUCUUGU ..........................((((((((((((...((.((((.....)))).)).....)).))))))))))......---....... ( -12.40) >DroSec_CAF1 15808 83 + 1 UUCAAUAAUUAUAGUUA-UUUUCUUGUAAUUUGUUAGGAGAAAAGUGUUUUUGGCACCUUGCGUGCCAUACCAAAUAUAUUCUU---------- .................-(((((((.(((....))).)))))))((((((.((((((.....))))))....))))))......---------- ( -17.20) >DroSim_CAF1 16274 92 + 1 UUCAAUAAUU--AGUUACUUUUCUUUUAAUUUGUUAGGAGAAAAGUGUUUUUGGCACCUUGUGUGCCAUACAAAACAUAUUCUUGUUACCUGGU .......(((--((..(((((((((((((....)))))))))))))(((((((((((.....))))))...))))).............))))) ( -23.30) >consensus UUCAAUAAUUAUAGUUACUUUUCUUGUAAUUUGUUAGGAGAAAAGUGUUUUUGGCACCUUGUGUGCCAUACCAAAUAUAUUCUU___A_CU_GU ...............((((((((((.(((....))).))))))))))....((((((.....)))))).......................... (-13.60 = -14.93 + 1.33)

| Location | 6,809,527 – 6,809,618 |
|---|---|
| Length | 91 |
| Sequences | 3 |
| Columns | 94 |
| Reading direction | reverse |
| Mean pairwise identity | 82.80 |
| Mean single sequence MFE | -16.21 |
| Consensus MFE | -12.81 |
| Energy contribution | -13.37 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.10 |
| Mean z-score | -2.61 |
| Structure conservation index | 0.79 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968131 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
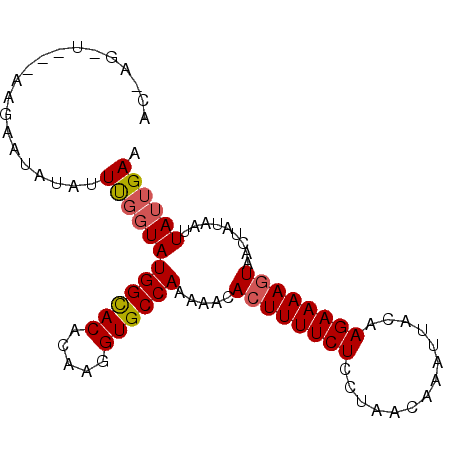
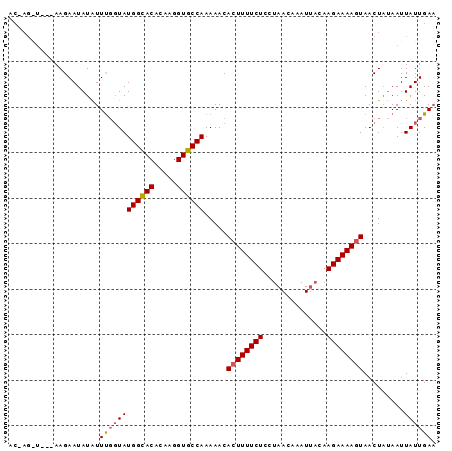
>2R_DroMel_CAF1 6809527 91 - 20766785 ACAAGAU---AAGAAUAUAUUCGUUAUGGUACACAAGGUGCCAAAAACACUUUUCUCCUAACAAAUAUCAAGAAAAGUAACUAUAAUUAUUGAU ....(((---((..(((.....(((.((((((.....))))))..)))((((((((..............))))))))...)))..)))))... ( -15.04) >DroSec_CAF1 15808 83 - 1 ----------AAGAAUAUAUUUGGUAUGGCACGCAAGGUGCCAAAAACACUUUUCUCCUAACAAAUUACAAGAAAA-UAACUAUAAUUAUUGAA ----------...((((....((((.((((((.....)))))).......((((((..(((....)))..))))))-..))))....))))... ( -14.60) >DroSim_CAF1 16274 92 - 1 ACCAGGUAACAAGAAUAUGUUUUGUAUGGCACACAAGGUGCCAAAAACACUUUUCUCCUAACAAAUUAAAAGAAAAGUAACU--AAUUAUUGAA ....(((((..((.....(((((...((((((.....)))))))))))((((((((..(((....)))..))))))))..))--..)))))... ( -19.00) >consensus AC_AG_U___AAGAAUAUAUUUGGUAUGGCACACAAGGUGCCAAAAACACUUUUCUCCUAACAAAUUACAAGAAAAGUAACUAUAAUUAUUGAA ....................((((((((((((.....)))))).....((((((((..............)))))))).........)))))). (-12.81 = -13.37 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:58 2006