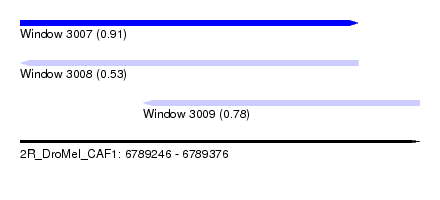

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,789,246 – 6,789,376 |
| Length | 130 |
| Max. P | 0.908688 |

| Location | 6,789,246 – 6,789,356 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | forward |
| Mean pairwise identity | 90.24 |
| Mean single sequence MFE | -17.82 |
| Consensus MFE | -15.46 |
| Energy contribution | -15.66 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.61 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.06 |
| SVM RNA-class probability | 0.908688 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6789246 110 + 20766785 CUCCACUUGGUGGCAACAUCCAAACCCCAACCCAUCCACAUCGAUCUGCCCCAAGCCCCUUCUCCUUUUU-UUUCGCUAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC ......((((.((((..(((......................))).))))))))((((.((((.......-...............)))).))))................ ( -17.30) >DroSec_CAF1 39393 108 + 1 CUCCACUUGGUGGCAACAUCCAAACCCCAAC--AUCCACAUCGAUCUGCCCCCAGCCCCUUCUCCUUUUU-UUUUGCUAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC ......((((((....)).))))....((((--(((......))).........((((.((((.......-...............)))).)))).))))........... ( -17.55) >DroSim_CAF1 48307 108 + 1 CUCCACUUGGUGGCAACAUCCAAACCCCAAC--AUCCACAUCGAUCUGCC-CCAGCCCCUUCUCCUUUUUUUUUUCCUAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC ......((((((....)).))))....((((--(((......))).....-...((((.((((.......................)))).)))).))))........... ( -17.50) >DroEre_CAF1 41970 103 + 1 CUCCACUUGGUGGCAACAUCCAAACCCCAAC--AACC-CAUCGAUCUGCC-GCAGCCCCUUCUCCAU----UUUCGUAAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC ......((((((....)).))))........--....-...(((...((.-((.((((.((((....----...............)))).)))).)).))......))). ( -19.51) >DroYak_CAF1 47175 97 + 1 CUCCACUUGGUGGCAACAUCC-------AAC--AUCC-CAUCGAUCUGCC-CCAGCCCCUUCUCCUUU---UUUCGUAAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC .......(((.((((..(((.-------...--....-....))).))))-)))((((.((((.....---...............)))).))))................ ( -17.25) >consensus CUCCACUUGGUGGCAACAUCCAAACCCCAAC__AUCCACAUCGAUCUGCC_CCAGCCCCUUCUCCUUUUU_UUUCGCUAACUUUUCAGAAAGGGCAGUUGCUUCUUUUCGC ........((.((((((................(((......))).........((((.((((.......................)))).)))).)))))).))...... (-15.46 = -15.66 + 0.20)

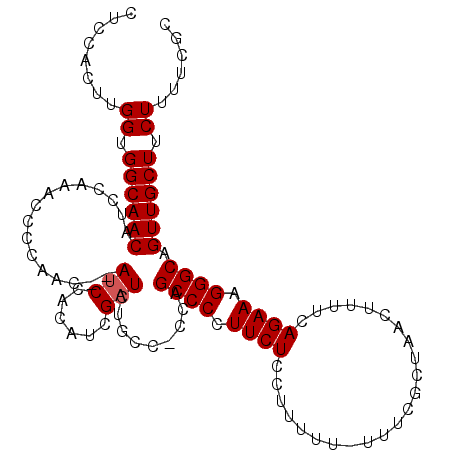
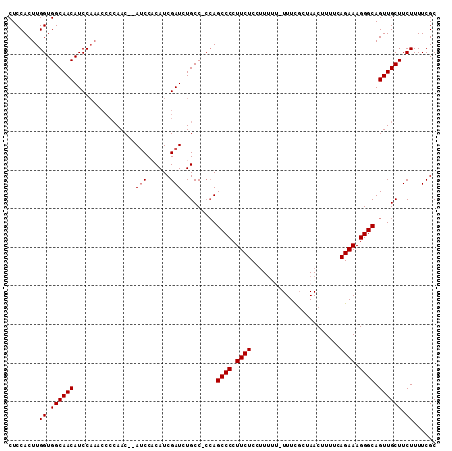
| Location | 6,789,246 – 6,789,356 |
|---|---|
| Length | 110 |
| Sequences | 5 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 90.24 |
| Mean single sequence MFE | -26.61 |
| Consensus MFE | -22.49 |
| Energy contribution | -22.89 |
| Covariance contribution | 0.40 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.28 |
| Structure conservation index | 0.85 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.525418 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6789246 110 - 20766785 GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCGAAA-AAAAAGGAGAAGGGGCUUGGGGCAGAUCGAUGUGGAUGGGUUGGGGUUUGGAUGUUGCCACCAAGUGGAG ((........))....(((((((((...............-.......)))))))))(((((((((((((((.(.((....)).).)))..))).))))).))))...... ( -27.45) >DroSec_CAF1 39393 108 - 1 GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCAAAA-AAAAAGGAGAAGGGGCUGGGGGCAGAUCGAUGUGGAU--GUUGGGGUUUGGAUGUUGCCACCAAGUGGAG ((........)).((((((((((((...............-.......)))))))))(((.(((((.(((((......--)))))..........))))).)))))).... ( -25.95) >DroSim_CAF1 48307 108 - 1 GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGGAAAAAAAAAAGGAGAAGGGGCUGG-GGCAGAUCGAUGUGGAU--GUUGGGGUUUGGAUGUUGCCACCAAGUGGAG ((........)).((((((((((((.......................)))))))))(((-(((((.(((((......--)))))..........))))).)))))).... ( -25.80) >DroEre_CAF1 41970 103 - 1 GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUUACGAAA----AUGGAGAAGGGGCUGC-GGCAGAUCGAUG-GGUU--GUUGGGGUUUGGAUGUUGCCACCAAGUGGAG .((((....((((((((((((((((.....((((....))----))..)))))))))..(-((....)))...-))))--)))....)))).......((((...)))).. ( -26.30) >DroYak_CAF1 47175 97 - 1 GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUUACGAAA---AAAGGAGAAGGGGCUGG-GGCAGAUCGAUG-GGAU--GUU-------GGAUGUUGCCACCAAGUGGAG ((........)).((((((((((((...............---.....)))))))))(((-(((((.(((((.-....--)))-------))...))))).)))))).... ( -27.55) >consensus GCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCGAAA_AAAAAGGAGAAGGGGCUGG_GGCAGAUCGAUGUGGAU__GUUGGGGUUUGGAUGUUGCCACCAAGUGGAG ((........)).((((((((((((.......................)))))))))(((.((((((((......................))).))))).)))))).... (-22.49 = -22.89 + 0.40)


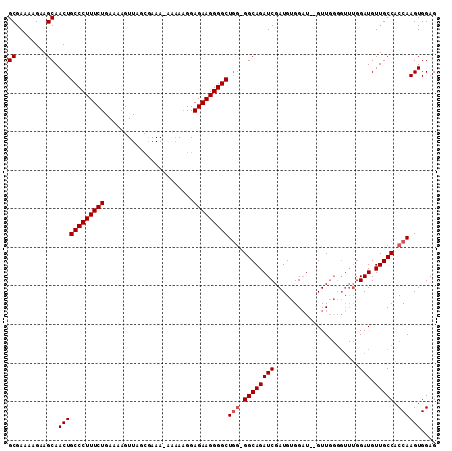
| Location | 6,789,286 – 6,789,376 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | reverse |
| Mean pairwise identity | 89.39 |
| Mean single sequence MFE | -22.47 |
| Consensus MFE | -16.10 |
| Energy contribution | -17.30 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.72 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778101 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6789286 90 - 20766785 AGUUUGUUUUCCCAAAUUCAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCGAAA-AAAAAGGAGAAGGGGCUUGGGGCAGAUCGA .((((((...(((((.....((........))....(((((((((...............-.......))))))))))))))))))))... ( -24.05) >DroSec_CAF1 39431 90 - 1 AGUUUGUUUUCCCAAAAGCAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCAAAA-AAAAAGGAGAAGGGGCUGGGGGCAGAUCGA .((((((..(((((......((........))....(((((((((...............-.......))))))))))))))))))))... ( -24.55) >DroSim_CAF1 48345 90 - 1 AGUUUGUUUUCCCAAAAGCAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGGAAAAAAAAAAGGAGAAGGGGCUGG-GGCAGAUCGA .((((((...((((......((........))....(((((((((.......................))))))))))))-)))))))... ( -23.10) >DroEre_CAF1 42007 82 - 1 AGU----UUUCCCAAAAGCAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUUACGAAA----AUGGAGAAGGGGCUGC-GGCAGAUCGA .((----(((....)))))..(((...(..(((...(((((((((.....((((....))----))..))))))))))))-..)...))). ( -19.30) >DroYak_CAF1 47205 82 - 1 AGU----UUUCCCAAAA-CAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUUACGAAA---AAAGGAGAAGGGGCUGG-GGCAGAUCGA .((----(((((((...-..((........))....(((((((((...............---.....))))))))))))-)).))))... ( -21.35) >consensus AGUUUGUUUUCCCAAAAGCAGCGAAAAGAAGCAACUGCCCUUUCUGAAAAGUUAGCGAAA_AAAAAGGAGAAGGGGCUGG_GGCAGAUCGA .((((.(((((.(.......).))))).))))....(((((((((.......................))))))))).............. (-16.10 = -17.30 + 1.20)


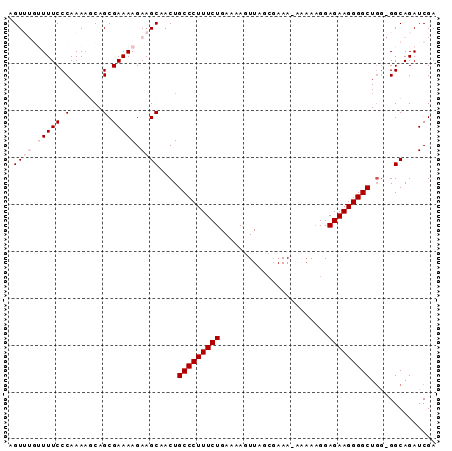
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:45 2006