
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,786,811 – 6,786,909 |
| Length | 98 |
| Max. P | 0.972424 |

| Location | 6,786,811 – 6,786,909 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | forward |
| Mean pairwise identity | 90.66 |
| Mean single sequence MFE | -33.80 |
| Consensus MFE | -24.53 |
| Energy contribution | -25.41 |
| Covariance contribution | 0.88 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.741827 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
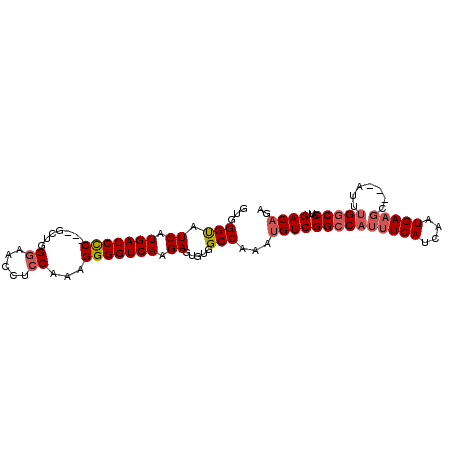
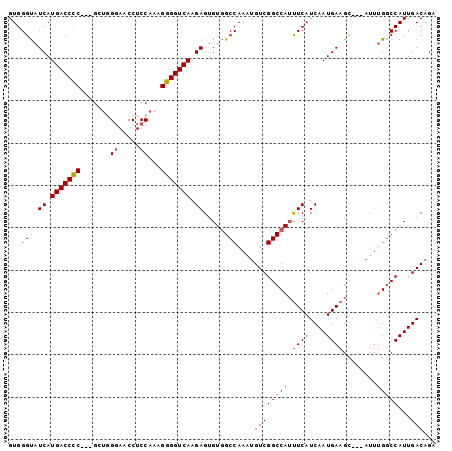
>2R_DroMel_CAF1 6786811 98 + 20766785 GUGGGUAUCAUGACCCCGCCGCUGGGAACCUCCAAAGGGGUCAAGAGUGUGGCCAAAUGUCGGCCAUUUCAUCAAUGAAGC---AUUUGUCCAUUGACAGA ....((.((((((((((.....((((....))))..))))))..((..((((((.......))))))....)).)))).))---.((((((....)))))) ( -33.10) >DroSec_CAF1 36961 95 + 1 GUGGGUAUCAUGACCCC---GCUGGGAACCUCCAAAGGGGUCAAGAGUGUGGCCAAAUGUCGGCCAUUUCAUCAAUGAAGC---AUAUGGCCAUUGACAGA ...(((.((.(((((((---..((((....))))..))))))).)).....)))...((((((((((((((....))))..---..))))))...)))).. ( -35.90) >DroSim_CAF1 45885 95 + 1 GUGGGUAUCAUGACCCC---GCUGGGAGCCUCCAAAGGGGUCAAGAGUGUGGCCAAAUGUCGGCCAUUUCAUCAAUGAAGC---AUUUGGCCAUUGACAGA .......((.(((((((---..((((....))))..))))))).))((((((((((((((....((((.....))))..))---))))))))))..))... ( -37.70) >DroEre_CAF1 39410 98 + 1 GUGGGUAUCAUGACCCC---GCCUGCAACCUCCAAAGGGGUCAAGAGUGUGGCCAAAUGUCGGCCAACUCAUCAAUGAAGACGAUGUUGGCCAUUGACUGA ...(((.((.(((((((---................))))))).)).....)))....((((((((((((.((......)).)).)))))))...)))... ( -33.29) >DroYak_CAF1 44702 95 + 1 GUGGGCAUCAUGACCUC---GCUCGGAACCUCCAAAGGGGUCAAGAGUGUAGCCAAAUGUCGGCCAUUUCAUCAAUGAAGC---AUUCGGCCAUUGACAGA ((((.(....(((((((---....((.....))...))))))).((((((.(((.......)))...((((....))))))---))))).))))....... ( -29.00) >consensus GUGGGUAUCAUGACCCC___GCUGGGAACCUCCAAAGGGGUCAAGAGUGUGGCCAAAUGUCGGCCAUUUCAUCAAUGAAGC___AUUUGGCCAUUGACAGA ...(((.((.(((((((.......((.....))...))))))).)).....)))...((((((((((((((....))))).......)))))...)))).. (-24.53 = -25.41 + 0.88)

| Location | 6,786,811 – 6,786,909 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 101 |
| Reading direction | reverse |
| Mean pairwise identity | 90.66 |
| Mean single sequence MFE | -34.60 |
| Consensus MFE | -26.24 |
| Energy contribution | -26.52 |
| Covariance contribution | 0.28 |
| Combinations/Pair | 1.07 |
| Mean z-score | -2.78 |
| Structure conservation index | 0.76 |
| SVM decision value | 1.69 |
| SVM RNA-class probability | 0.972424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
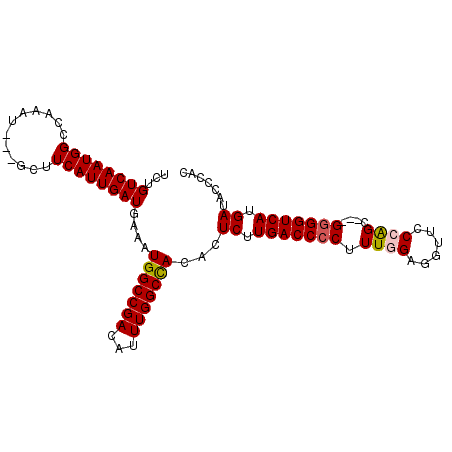
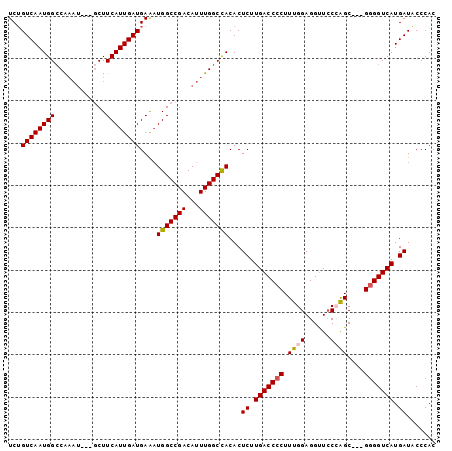
>2R_DroMel_CAF1 6786811 98 - 20766785 UCUGUCAAUGGACAAAU---GCUUCAUUGAUGAAAUGGCCGACAUUUGGCCACACUCUUGACCCCUUUGGAGGUUCCCAGCGGCGGGGUCAUGAUACCCAC ...((((((((((....---).)))))))))....(((((((...)))))))...((.(((((((.((((......))))....))))))).))....... ( -32.90) >DroSec_CAF1 36961 95 - 1 UCUGUCAAUGGCCAUAU---GCUUCAUUGAUGAAAUGGCCGACAUUUGGCCACACUCUUGACCCCUUUGGAGGUUCCCAGC---GGGGUCAUGAUACCCAC ..(((((.((((((.((---((.(((((.....)))))..).))).))))))......(((((((.((((......)))).---))))))))))))..... ( -35.10) >DroSim_CAF1 45885 95 - 1 UCUGUCAAUGGCCAAAU---GCUUCAUUGAUGAAAUGGCCGACAUUUGGCCACACUCUUGACCCCUUUGGAGGCUCCCAGC---GGGGUCAUGAUACCCAC ..(((((.(((((((((---((.(((((.....)))))..).))))))))))......(((((((.((((......)))).---))))))))))))..... ( -38.40) >DroEre_CAF1 39410 98 - 1 UCAGUCAAUGGCCAACAUCGUCUUCAUUGAUGAGUUGGCCGACAUUUGGCCACACUCUUGACCCCUUUGGAGGUUGCAGGC---GGGGUCAUGAUACCCAC ...(((((((((((((.(((((......)))))))))))))....))))).....((.(((((((((((.......)))).---))))))).))....... ( -37.60) >DroYak_CAF1 44702 95 - 1 UCUGUCAAUGGCCGAAU---GCUUCAUUGAUGAAAUGGCCGACAUUUGGCUACACUCUUGACCCCUUUGGAGGUUCCGAGC---GAGGUCAUGAUGCCCAC ...((((.(((((((((---((.(((((.....)))))..).))))))))))......((((((.(((((.....))))).---).)))))))))...... ( -29.00) >consensus UCUGUCAAUGGCCAAAU___GCUUCAUUGAUGAAAUGGCCGACAUUUGGCCACACUCUUGACCCCUUUGGAGGUUCCCAGC___GGGGUCAUGAUACCCAC ...((((((((............))))))))....(((((((...)))))))...((.(((((((.((((......))))....))))))).))....... (-26.24 = -26.52 + 0.28)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:37 2006