
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,718,209 – 6,718,300 |
| Length | 91 |
| Max. P | 0.794022 |

| Location | 6,718,209 – 6,718,300 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 89.33 |
| Mean single sequence MFE | -17.52 |
| Consensus MFE | -12.80 |
| Energy contribution | -12.76 |
| Covariance contribution | -0.04 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.45 |
| SVM RNA-class probability | 0.740424 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
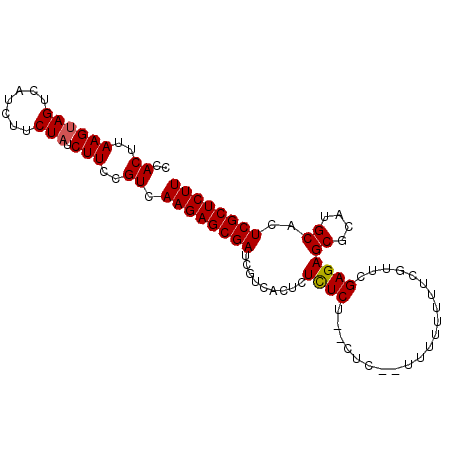
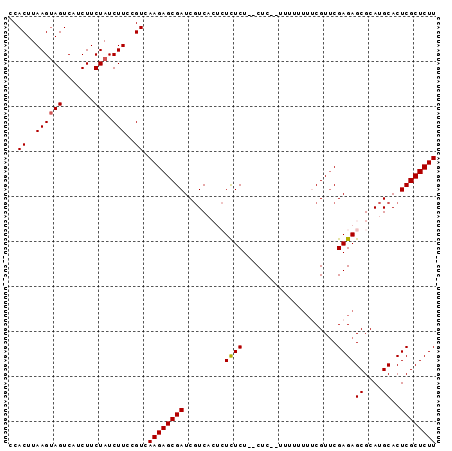
>2R_DroMel_CAF1 6718209 91 + 20766785 CCACUUAAGUAGUCAUCUUCUAUCUUCCGUCAAGAGCGAUCGUCACUCUCUCU--CUC--UUUUUUUUCGUUCGAGAGCGCAUGCACUCGCUCUU ..((..((((((.......))).)))..)).((((((((.((((.(((((...--...--.............))))).).)))...)))))))) ( -17.81) >DroSec_CAF1 66435 93 + 1 CCACUUAAGUAGUCAUCUUCUAUCUUCCGUCAAGAGCGAUCGUCACUCUCUCU--CUCUUUUUAUUUUCGUUCGAGAGCGCAUGCACUCGCUCUU ..((..((((((.......))).)))..)).((((((((.((((.(((((...--..................))))).).)))...)))))))) ( -17.70) >DroSim_CAF1 74185 95 + 1 CCACUUAAGCAGUCAUCUUCUAUCUUCCGUCAAGAGCGAUCGUCACUCUCUCUCUCUCUCUUUUUUUUCGUUCGAGAGCGCAUGCACUCGCUCUU ...............................((((((((.((((.(((((.......................))))).).)))...)))))))) ( -16.90) >DroEre_CAF1 68939 82 + 1 CCACUUAAGUAGUCAUCUUCUAUCUUCUGUCAAGAGCGAUC-UC----UUUCU--CUC--UUUU----CGUACGAGAGCGCAUGCACUCGCUCUU ..((..((((((.......))).)))..)).((((((((..-..----...((--(((--....----.....))))).........)))))))) ( -16.39) >DroYak_CAF1 67416 87 + 1 CCACUUAAGUAGUCAUCUUCUAUCUUCGGUCAAGAGCGAUCGUC----UUUCU--CUC--UUUCCUUGCGUUCGAGAGCGCAUGCACUCGCUCUU ..(((.((((((.......))).))).))).((((((((..((.----...((--(((--.............))))).....))..)))))))) ( -18.82) >consensus CCACUUAAGUAGUCAUCUUCUAUCUUCCGUCAAGAGCGAUCGUCACUCUCUCU__CUC__UUUUUUUUCGUUCGAGAGCGCAUGCACUCGCUCUU ..((..((((((.......))).)))..)).((((((((.........((((.....................))))((....))..)))))))) (-12.80 = -12.76 + -0.04)

| Location | 6,718,209 – 6,718,300 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 89.33 |
| Mean single sequence MFE | -24.06 |
| Consensus MFE | -16.52 |
| Energy contribution | -17.96 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.04 |
| Mean z-score | -2.69 |
| Structure conservation index | 0.69 |
| SVM decision value | 0.60 |
| SVM RNA-class probability | 0.794022 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
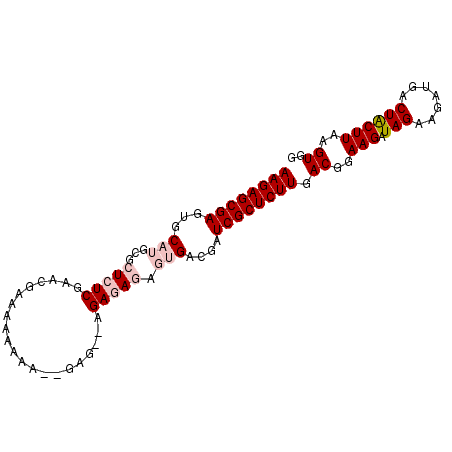
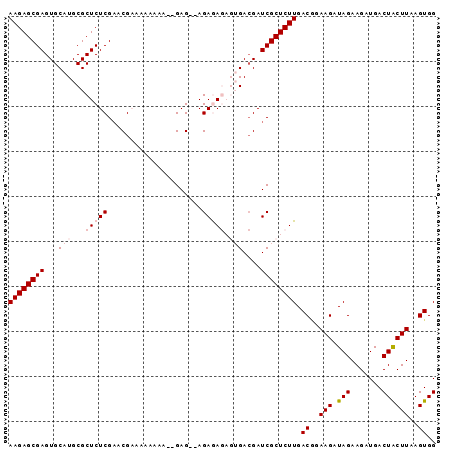
>2R_DroMel_CAF1 6718209 91 - 20766785 AAGAGCGAGUGCAUGCGCUCUCGAACGAAAAAAAA--GAG--AGAGAGAGUGACGAUCGCUCUUGACGGAAGAUAGAAGAUGACUACUUAAGUGG ((((((((.(((((...(((((.............--..)--))))...))).)).))))))))..(....)...........((((....)))) ( -25.56) >DroSec_CAF1 66435 93 - 1 AAGAGCGAGUGCAUGCGCUCUCGAACGAAAAUAAAAAGAG--AGAGAGAGUGACGAUCGCUCUUGACGGAAGAUAGAAGAUGACUACUUAAGUGG ((((((((.(((((...(((((.................)--))))...))).)).))))))))..(....)...........((((....)))) ( -25.43) >DroSim_CAF1 74185 95 - 1 AAGAGCGAGUGCAUGCGCUCUCGAACGAAAAAAAAGAGAGAGAGAGAGAGUGACGAUCGCUCUUGACGGAAGAUAGAAGAUGACUGCUUAAGUGG ((((((((.(((((...(((((...................)))))...))).)).)))))))).((..(((.(((.......))))))..)).. ( -25.01) >DroEre_CAF1 68939 82 - 1 AAGAGCGAGUGCAUGCGCUCUCGUACG----AAAA--GAG--AGAAA----GA-GAUCGCUCUUGACAGAAGAUAGAAGAUGACUACUUAAGUGG ((((((((.........(((((.....----....--)))--))...----..-..))))))))...................((((....)))) ( -20.09) >DroYak_CAF1 67416 87 - 1 AAGAGCGAGUGCAUGCGCUCUCGAACGCAAGGAAA--GAG--AGAAA----GACGAUCGCUCUUGACCGAAGAUAGAAGAUGACUACUUAAGUGG ((((((((.((..(...(((((...(....)....--)))--))..)----..)).)))))))).((..(((.(((.......))))))..)).. ( -24.20) >consensus AAGAGCGAGUGCAUGCGCUCUCGAACGAAAAAAAA__GAG__AGAGAGAGUGACGAUCGCUCUUGACGGAAGAUAGAAGAUGACUACUUAAGUGG ((((((((...(((...(((((.....................))))).)))....)))))))).((..(((.(((.......))))))..)).. (-16.52 = -17.96 + 1.44)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:22 2006