
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,710,257 – 6,710,348 |
| Length | 91 |
| Max. P | 0.600289 |

| Location | 6,710,257 – 6,710,348 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 75.85 |
| Mean single sequence MFE | -36.60 |
| Consensus MFE | -25.73 |
| Energy contribution | -26.85 |
| Covariance contribution | 1.12 |
| Combinations/Pair | 1.16 |
| Mean z-score | -1.48 |
| Structure conservation index | 0.70 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.600289 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
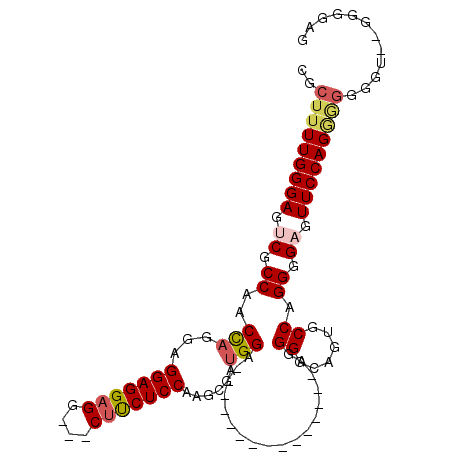
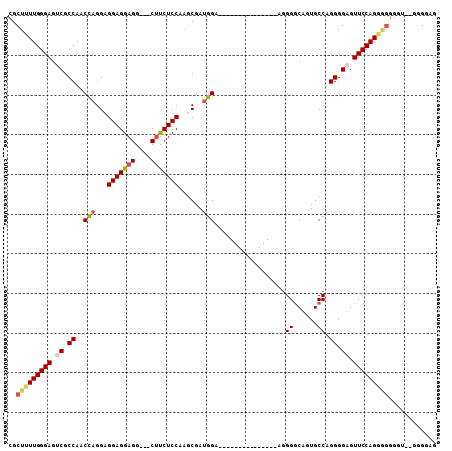
>2R_DroMel_CAF1 6710257 91 - 20766785 CGCUUUUGGGAGUCGCCAACCAGGAGGAGGAGGUUGCCUCUCCAAGCGAUGGA---------------AGGGGCUGUGCCAGGGGAGUUCCAGAAGGGGU--GAGUAG (.(((((((((.((.((..((....))....(((.(((((((((.....))).---------------))))))...))).)).)).))))))))).)..--...... ( -38.80) >DroSec_CAF1 58724 86 - 1 CGCUUUUGGGAGUCGCCAACCAGGCGGAGGAGG---CUUCUCCAAGCGAUGGA---------------AGGGGCAGUGCCAGGGGUGUUCCAGGGGGU----GGGGAG ((((((..((((.((((..((.((((......(---((((((((.....))).---------------))))))..)))).))))))))))..)))))----)..... ( -32.40) >DroSim_CAF1 65525 88 - 1 CGCUUUUGGGAGUCGCCAACCAGGCGGAGGAGG---CUUCUCCAAGCGAUGGA---------------ACGGGCAGUGCCAGGGGUGUUCCAGGGGGUUU--GGGGGG .((((((.....(((((.....))))).)))))---).(((((((((..((((---------------((((((...))).....)))))))....))))--))))). ( -33.90) >DroYak_CAF1 59562 100 - 1 CGCUUUUGGGAGUCGCCAACU-GGAGGAGGAGG---CUCCUCCACACGAUGGGGGUGCUCAGGGGCGGAGGGACUGUGCCAGGGGAGUUCCAG----GGGAAGGGGGU ((((((((((..((.(((.((-(((((((....---))))))))...).))).))..))))))))))..((((((.(......).))))))..----........... ( -41.30) >consensus CGCUUUUGGGAGUCGCCAACCAGGAGGAGGAGG___CUUCUCCAAGCGAUGGA_______________AGGGGCAGUGCCAGGGGAGUUCCAGGGGGGGU__GGGGAG ..(((((((((.((.((..(((...(((((((....)))))))......)))..................((......)).)).)).)))))))))............ (-25.73 = -26.85 + 1.12)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:15 2006