
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,680,157 – 6,680,251 |
| Length | 94 |
| Max. P | 0.607422 |

| Location | 6,680,157 – 6,680,251 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 77.88 |
| Mean single sequence MFE | -15.92 |
| Consensus MFE | -2.12 |
| Energy contribution | -3.28 |
| Covariance contribution | 1.16 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.63 |
| Structure conservation index | 0.13 |
| SVM decision value | 0.15 |
| SVM RNA-class probability | 0.607422 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
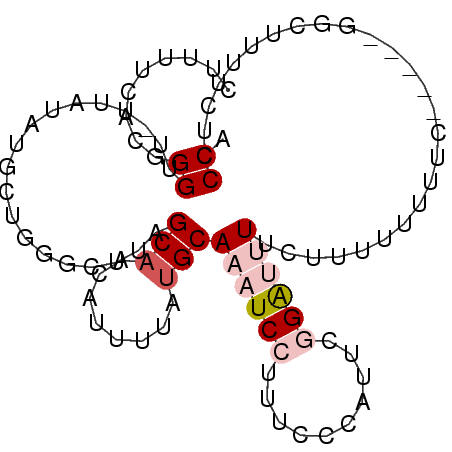
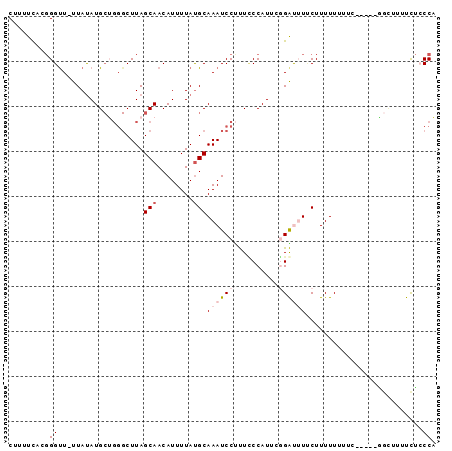
>2R_DroMel_CAF1 6680157 94 - 20766785 CUUUUCACGGGCU-UUAUAUGCUGGGCUUAGCAACAUUUUAUGCAAAUCCGUUCCCAUUCGGAUUUGCUUUUUUUUUUUGGGGGGUUUUCUCCCA .......(.(((.-......))).).................(((((((((........)))))))))............(((((....))))). ( -25.80) >DroSec_CAF1 28651 89 - 1 CUUUUUAGGGGUU-UUAUAUGCUGGGCUUAGCAACAUUUUAUGCAAAUCCUUUUCCAUUCGGAUUUCCUUUUUUUUC-----AGCUUGUCUACCA .........(((.-..(((.((((((....(((........)))((((((..........))))))........)))-----))).)))..))). ( -16.20) >DroSim_CAF1 31708 89 - 1 CUUUUCACGGGUU-UUAUAUGCUGGGCUUAGCAACAUUUUAUGCAAAUCCUUUUCCAUUCGGAUUUACUUUUUUUUC-----GGCUUGUCUACCA .........(((.-..(((.((((((....(((........)))((((((..........))))))........)))-----))).)))..))). ( -16.10) >DroEre_CAF1 26329 84 - 1 CUUUUCACGGGUUUUUACAUG-----CUUAGCAACAUUUUACGCAAAUCCCCUACCAUUCUGGCAUUCUUUUU-GGC-----UGUUUUUUUCCCA ........(((.....(((.(-----((..((..........))..........((.....))..........-)))-----)))......))). ( -10.20) >DroYak_CAF1 28603 85 - 1 CUUUUCAUCGGUU-UUAUAUGCGGGGCUUAGCAACAUUUUAUGCAAAUCCUCUCCCAUUCCGGCAUUCUUUUU-GGC-----U---UUUUACCCU .........(((.-......((((((.((.(((........))).)).))))..((.....))..........-.))-----.---....))).. ( -11.30) >consensus CUUUUCACGGGUU_UUAUAUGCUGGGCUUAGCAACAUUUUAUGCAAAUCCUUUCCCAUUCGGAUUUUCUUUUUUUUC_____GGCUUUUCUCCCA .........((...................(((........)))((((((..........))))))..........................)). ( -2.12 = -3.28 + 1.16)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:13:03 2006