| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,568,003 – 6,568,094 |
| Length | 91 |
| Max. P | 0.960034 |

| Location | 6,568,003 – 6,568,094 |
|---|---|
| Length | 91 |
| Sequences | 4 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 77.59 |
| Mean single sequence MFE | -25.09 |
| Consensus MFE | -20.66 |
| Energy contribution | -20.48 |
| Covariance contribution | -0.19 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.89 |
| Structure conservation index | 0.82 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.960034 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6568003 91 - 20766785 UGUCAGCCCGGCCGGGAACCCUUUCAUGGGCACAAAAACCAGCCAAAGGGGUGUGACCCAGACCCACAUCU-----CCCACACCCAUA-----CCCCAUAC----------- .....(((((...((....)).....)))))................((((((((................-----........))))-----))))....----------- ( -23.56) >DroSec_CAF1 120568 104 - 1 UGUCAGCCCGGCCGGGAACCCUUUAACGGGCACAAAAACCAGCCAAAGGGGUGUGACCCAGACCCACACCC--------GCACCCACACCAUUCCCCAACCCCCCUCCACCA .....(((((...((....)).....))))).................(((((((.........)))))))--------................................. ( -24.70) >DroEre_CAF1 121096 101 - 1 UGUCAGCCCGGCCGGGAACCCUUUCACGGGCACAAAAACCACCCAAAGGGGUGUGACCCAGACCCAUACCCCCUAUCCCCUACCCACACCCAUGGCCACAC----------- .........(((((((..(((......))).................((((((((.........))))))))................)))..))))....----------- ( -29.00) >DroYak_CAF1 126563 103 - 1 UGUCAGCCCGGCCGGGAACCCUUUCACGGGCACAAAAACCAGCCAAAGGGGUGUGACCCAGACCCAGACUCA--UUCCCAUACCAACACCAUUCCCCAGAACU-------GA .(((.(((((...((....)).....))))).................((((.((...)).)))).)))...--.............................-------.. ( -23.10) >consensus UGUCAGCCCGGCCGGGAACCCUUUCACGGGCACAAAAACCAGCCAAAGGGGUGUGACCCAGACCCACACCC_____CCCACACCCACACCAUUCCCCAAAC___________ .(((.(((((...((....)).....)))))................(((......))).)))................................................. (-20.66 = -20.48 + -0.19)

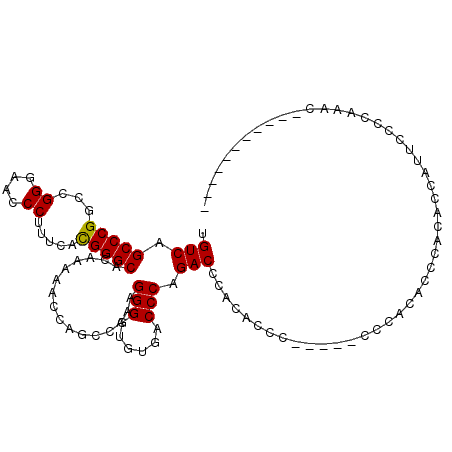
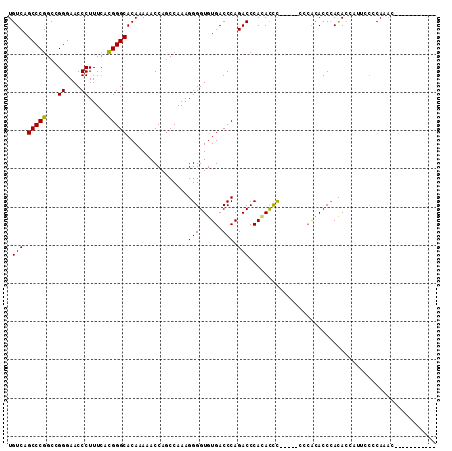
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:12:15 2006