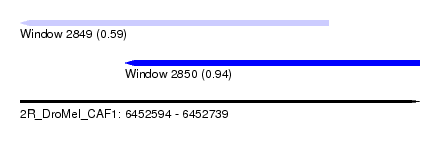
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,452,594 – 6,452,739 |
| Length | 145 |
| Max. P | 0.942237 |

| Location | 6,452,594 – 6,452,706 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 83.19 |
| Mean single sequence MFE | -21.05 |
| Consensus MFE | -15.44 |
| Energy contribution | -15.50 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.73 |
| SVM decision value | 0.11 |
| SVM RNA-class probability | 0.589221 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
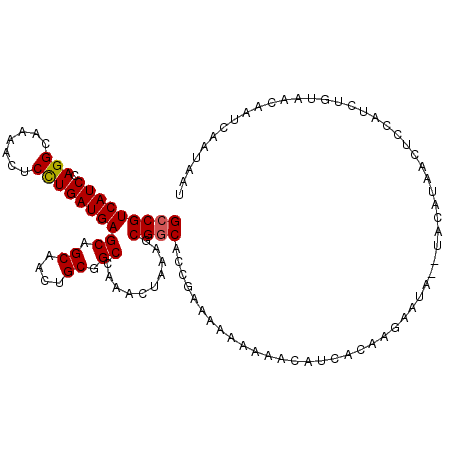
>2R_DroMel_CAF1 6452594 112 - 20766785 GCCGUCAUCCAGGCAAAACUCUUGAUGAGCAGCAACUGCGGCCAAACUAAAGCGACACCGAAAAAAAAACAACACAAGAAUA--UACAUCACUCUAUCUUUAAGAAACAAUAAU (((.(((((.(((......))).)))))((((...)))))))........................................--.............................. ( -14.50) >DroSec_CAF1 28319 110 - 1 GCCGUCAUCCAGGCAAAACUCCUGAUGAGCAGCAACUGCGGCCAAACUAAAGCGGCACCGAAAA--AAACAGCACAAGGAUA--UACAACACUCCAUCUUUAAGAAUCAAUAAU (((.(((((.(((.......))))))))((((...))))))).....(((((((....))....--...........(((..--........)))..)))))............ ( -19.40) >DroEre_CAF1 26864 114 - 1 GCCGUCAUCCAGGCAAAACUCCUGAUGAGCGGCAACUGCGGCCAAACUAAAGCGGCACCGCAAAAAAUGCCUCACAAGAAUACAUAUGUACCUUCAUCCGUACCUAUCAAUUAU (((((((((.(((.......)))))))).))))...(((((..........(((....)))................((((((....)))..)))..)))))............ ( -26.80) >DroYak_CAF1 30047 112 - 1 GCCGUCAUCCAGGCAAAACUCCUGAUGAGCUGCAACUGCGGCCAAACUAAAGCGGCACCGAAAAAAAAAACUCUCAAUAAUA--UAUGCACCUUAAUAUGUAACGAUCAAUAAG (((((((((.(((.......))))))))(((((....)))))..........)))).......................(((--(((........))))))............. ( -23.50) >consensus GCCGUCAUCCAGGCAAAACUCCUGAUGAGCAGCAACUGCGGCCAAACUAAAGCGGCACCGAAAAAAAAACAUCACAAGAAUA__UACAUAACUCCAUCUGUAACAAUCAAUAAU (((((((((.(((.......))))))))((.((....)).))..........)))).......................................................... (-15.44 = -15.50 + 0.06)

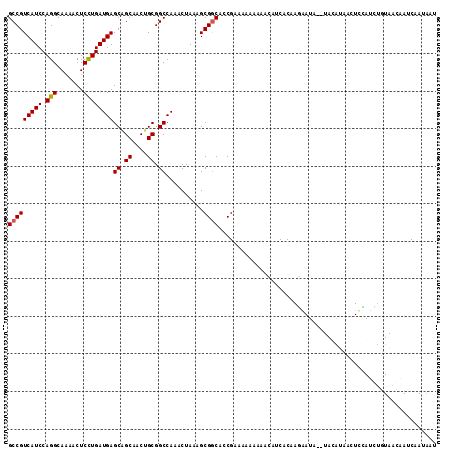
| Location | 6,452,632 – 6,452,739 |
|---|---|
| Length | 107 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | reverse |
| Mean pairwise identity | 85.93 |
| Mean single sequence MFE | -32.60 |
| Consensus MFE | -25.96 |
| Energy contribution | -26.68 |
| Covariance contribution | 0.72 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.17 |
| Structure conservation index | 0.80 |
| SVM decision value | 1.32 |
| SVM RNA-class probability | 0.942237 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
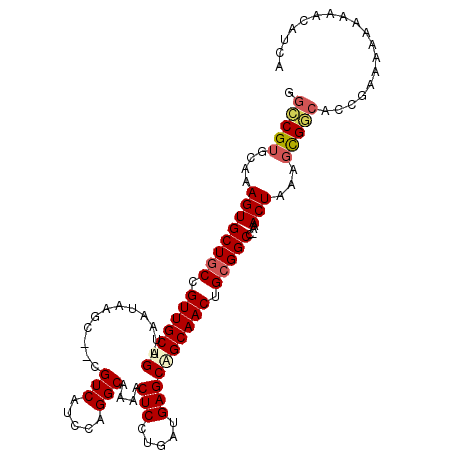
>2R_DroMel_CAF1 6452632 107 - 20766785 GGCCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGC--CGUCAUCCAGGCAAAACUCUUGAUGAGCAGCAACUGCGGCC-AAACUAAAGCGACACCGAAAAAAAAACAACA ((...(((..((((((((.(((((((.......((--(........)))....(((.....)))))))))).))))).-..)))...)))...))............... ( -32.20) >DroSec_CAF1 28357 105 - 1 GGCCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGC--CGUCAUCCAGGCAAAACUCCUGAUGAGCAGCAACUGCGGCC-AAACUAAAGCGGCACCGAAAA--AAACAGCA .(((((....((((((((.(((((((.......((--(........)))....(((.....)))))))))).))))).-..)))...)))))........--........ ( -36.10) >DroEre_CAF1 26904 107 - 1 GGCCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGC--CGUCAUCCAGGCAAAACUCCUGAUGAGCGGCAACUGCGGCC-AAACUAAAGCGGCACCGCAAAAAAUGCCUCA .(((((....((((((((.(((((((.......((--(........)))....(((.....)))))))))).))))).-..)))...)))))...(((.....))).... ( -36.70) >DroYak_CAF1 30085 107 - 1 GGCCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGC--CGUCAUCCAGGCAAAACUCCUGAUGAGCUGCAACUGCGGCC-AAACUAAAGCGGCACCGAAAAAAAAAACUCU .(((((....((((((((.(((((........(((--..(((((.(((.......)))))))))))))))).))))).-..)))...))))).................. ( -32.70) >DroAna_CAF1 24989 99 - 1 UGGCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGCCCCGUCAUCCCGGCAAAACUCCUGAUGAGCUGCAACAGAGGCUGAAACUGAGUUGCAACCGAAG----------- .((((.((.....))))))(((((.........(((..........)))..(((((....(.((((........)))).)....)))))))))).....----------- ( -25.30) >consensus GGCCGUGCAAAGUGCUGCCGUUGCUGAUAAUAAGC__CGUCAUCCAGGCAAAACUCCUGAUGAGCAGCAACUGCGGCC_AAACUAAAGCGGCACCGAAAAAAAAACAUCA .(((((....((((((((.(((((((............(((.....)))....(((.....)))))))))).)))))....)))...))))).................. (-25.96 = -26.68 + 0.72)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:11:15 2006