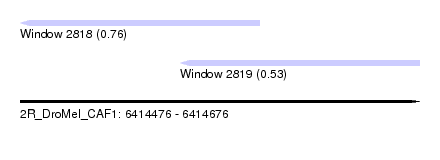
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,414,476 – 6,414,676 |
| Length | 200 |
| Max. P | 0.762885 |

| Location | 6,414,476 – 6,414,596 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 94.42 |
| Mean single sequence MFE | -41.76 |
| Consensus MFE | -38.90 |
| Energy contribution | -39.10 |
| Covariance contribution | 0.20 |
| Combinations/Pair | 1.06 |
| Mean z-score | -1.20 |
| Structure conservation index | 0.93 |
| SVM decision value | 0.51 |
| SVM RNA-class probability | 0.762885 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
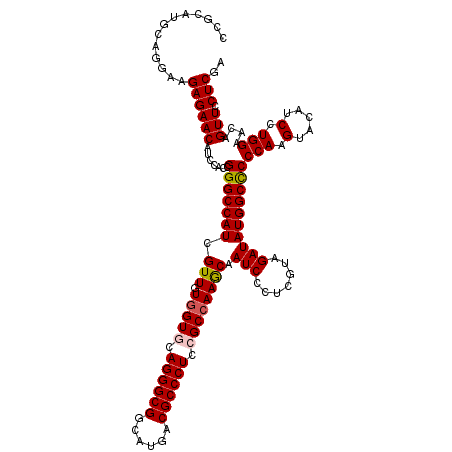
>2R_DroMel_CAF1 6414476 120 - 20766785 UCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUGGUGCAGGGCGGCAUGACGCCCUCCGCCAAGCAAUCGCUCGUAGAUAUGGCCCCCAAGUACAUCCUGGAACAGUUUCUCGA .............((((((.......(((((((.(((.(((((.((((((......)))))).)))))))).(((.......))))))))))(((.(.....).)))....).))))).. ( -42.00) >DroSec_CAF1 10222 120 - 1 CCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUGGUGCAGGGCGGCAUGACGCCCUCCGCCAAGCAAUCCCUCGUAGAUAUGGCCCCCAAGUACAUCCUGGAACAGUUCCUCGA ((.......))..((((((.......(((((((.(((.(((((.((((((......)))))).)))))))).(((.......))))))))))(((.(.....).)))....))).))).. ( -41.50) >DroSim_CAF1 12316 120 - 1 CCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUGGUGCAGGGCGGCAUGACGCCCUCCGCCAAGCAAUCCCUCGUAGAUAUGGCCCCCAAGUACAUCCUGGAACAGUUCCUCGA ((.......))..((((((.......(((((((.(((.(((((.((((((......)))))).)))))))).(((.......))))))))))(((.(.....).)))....))).))).. ( -41.50) >DroEre_CAF1 8983 120 - 1 CCGUAUGCAAGAAGAGAACAUCCACCGGGCCAUCGUUGUUGUUCAGGGCGGAAUGACGCCCUCCGCCAAACAAUCCCUCGUGGAUAUGGCUCCCAAGUACAUCCUGGAGCAGUUCCUCGA .............((((((((((((.(((.....((((((((..((((((......))))))..))..)))))))))..))))))...(((((..((......))))))).))).))).. ( -39.90) >DroYak_CAF1 9135 120 - 1 CCGUAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUGGUUCAGGGCGGAAUGACGCCCUCCGCCAAACAAUCUCUCGUGGAUAUGGCCCCCAAGUACAUCUUGGAGCAGUUCCUCGA ((.......))..((((((.......(((((((.(((.((((..((((((......))))))..))))))).((((.....)))))))))))(((((.....)))))....))).))).. ( -43.90) >consensus CCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUGGUGCAGGGCGGCAUGACGCCCUCCGCCAAGCAAUCCCUCGUAGAUAUGGCCCCCAAGUACAUCCUGGAACAGUUCCUCGA .............((((((.......(((((((.(((.(((((.((((((......)))))).)))))))).(((.......))))))))))(((.(.....).)))....))).))).. (-38.90 = -39.10 + 0.20)

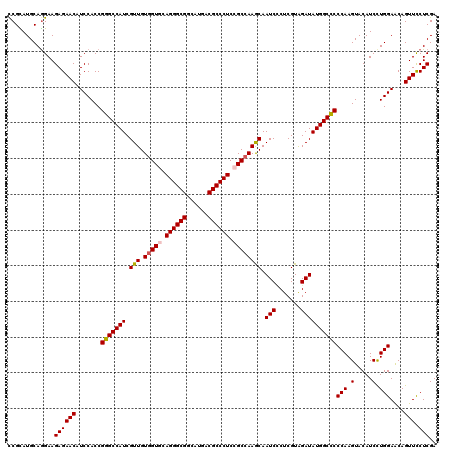
| Location | 6,414,556 – 6,414,676 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 96.67 |
| Mean single sequence MFE | -38.00 |
| Consensus MFE | -36.96 |
| Energy contribution | -36.52 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.05 |
| Structure conservation index | 0.97 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.526918 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6414556 120 - 20766785 GACGAUCCCACGGACCAGAUGUUCGUCUUCUUUCCCGAGGAGCCGAAGAUCGGCAUCAAGACAAUCAAGACGUACUGCACUCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUG ((((((.((.(((....(((((((.(((((.........((((((.....)))).)).................(((((......)))))))))))))))))..)))))..))))))... ( -39.80) >DroSec_CAF1 10302 120 - 1 GACGAUCCCACGGACCAGAUGUUCGUCUUCUUUCCCGAGGAGCCGAAGAUCGGCAUCAAGACAAUCAAGACGUACUGCACCCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUG ((((((.((.(((....(((((((.(((((.........((((((.....)))).)).................(((((......)))))))))))))))))..)))))..))))))... ( -39.80) >DroSim_CAF1 12396 120 - 1 GACGAUCCCACGGACCAGAUGUUCGUCUUCUUUCCCGAGGAGCCGAAGAUCGGCAUCAAGACAAUCAAGACGUACUGCACCCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUG ((((((.((.(((....(((((((.(((((.........((((((.....)))).)).................(((((......)))))))))))))))))..)))))..))))))... ( -39.80) >DroEre_CAF1 9063 120 - 1 GACGAUCCCACGGACCAGAUGUUUGUCUUCUUCCCCGAGGAGCCGAAGAUCGGCAUCAAGACCAUCAAGACGUACUGCACCCGUAUGCAAGAAGAGAACAUCCACCGGGCCAUCGUUGUU ((((((.((.(((....(((((((.(((((((.......((((((.....)))).))...........(.(((((.......))))))))))))))))))))..)))))..))))))... ( -36.00) >DroYak_CAF1 9215 120 - 1 GACGAUCCCACGGACCAGAUGUUUGUCUUCUUCCCCGAGGAGCCAAAGAUCGGCAUCAAGACCAUCAAGACGUACUGCACCCGUAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUG ((((((.((.(((....(((((((.(((((.(((....)))(((.......)))....................(((((......)))))))))))))))))..)))))..))))))... ( -34.60) >consensus GACGAUCCCACGGACCAGAUGUUCGUCUUCUUUCCCGAGGAGCCGAAGAUCGGCAUCAAGACAAUCAAGACGUACUGCACCCGCAUGCAGGAAGAGAACAUCCACCGGGCCAUCGUUGUG ((((((.((.(((....(((((((.(((((((.......((((((.....)))).))..............(((.(((....))))))))))))))))))))..)))))..))))))... (-36.96 = -36.52 + -0.44)

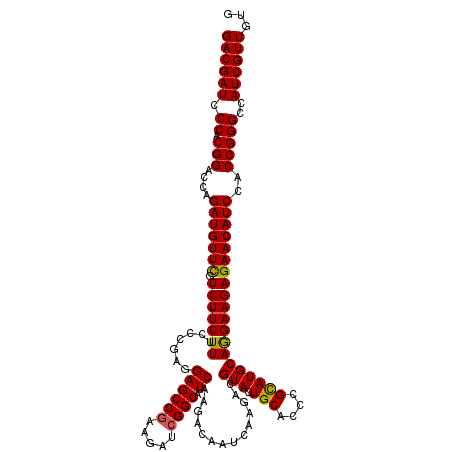
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:46 2006