
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,378,962 – 6,379,077 |
| Length | 115 |
| Max. P | 0.711592 |

| Location | 6,378,962 – 6,379,077 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | forward |
| Mean pairwise identity | 83.46 |
| Mean single sequence MFE | -30.40 |
| Consensus MFE | -16.06 |
| Energy contribution | -17.00 |
| Covariance contribution | 0.94 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.63 |
| Structure conservation index | 0.53 |
| SVM decision value | 0.13 |
| SVM RNA-class probability | 0.599748 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6378962 115 + 20766785 ACAUGUGUGGGGCAUCUGUAUCCAUGCCCAAGAACUGUCUGUAAGCUAUUUAUGAACCCAUCAAGUUCAUGUUCAUCGAAAACGAGCAAUGAUCUAUAGUCAAUACACAUGUC--UA (((((((((((((((........))))))......((.(((((........((((((.......))))))((((.........)))).......))))).)).))))))))).--.. ( -31.30) >DroSec_CAF1 19625 115 + 1 ACAUGUGUGGGGCAUCUGUAUCCAUGCCCAAGAACUGUCUGUAAGCUAUUUAUGAACCCGUCAAGUUCAUGUUCAUCGCAAACGAGUAAUGGUCUAUAGACAAUAGACAUGUC--UA ((((((.((((((((........))))))......((((((((..((((((((((((.......)))))))....(((....)))..)))))..)))))))).)).)))))).--.. ( -36.50) >DroEre_CAF1 20836 102 + 1 ACAUGUGUGGGGCAUGUGUAUC----CCCAAGAAGUG----UAAGCUAUUUAUGAACUCAUCAAGUUCAUGUUCAUCGAAACCAAACAAUGAUCUAUAGAC-----ACAUGCC--CG .........((((((((((...----....((((((.----...)))...((((((((.....)))))))).....................)))....))-----)))))))--). ( -26.60) >DroYak_CAF1 20898 104 + 1 ACAUGUGUGGGGCAUGUGUAUC----CCCAAGAAGUG----UCAGCUAUUUAUGAACUCAUCAAGUUCAUAUUCAUCGAAACCGAGCAAUGAUCUAUAGGC-----ACAUGUCUCUA ((((((((((((.((....)))----)))....((.(----(((((....((((((((.....))))))))....(((....)))))..))))))....))-----))))))..... ( -27.20) >consensus ACAUGUGUGGGGCAUCUGUAUC____CCCAAGAACUG____UAAGCUAUUUAUGAACCCAUCAAGUUCAUGUUCAUCGAAAACGAGCAAUGAUCUAUAGAC_____ACAUGUC__UA .(((((...((((((........)))))).(((.................(((((((.......))))))).((((.(........).)))))))...........)))))...... (-16.06 = -17.00 + 0.94)

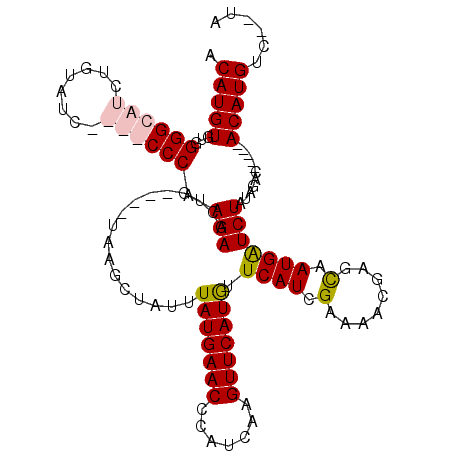
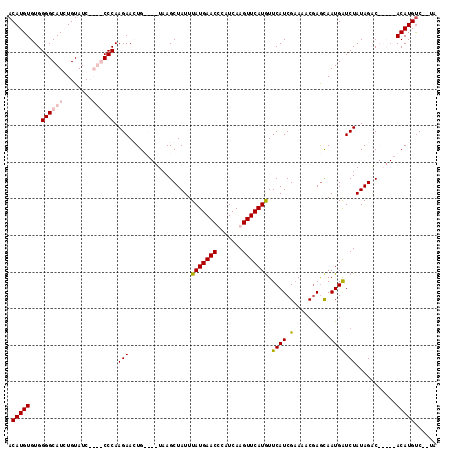
| Location | 6,378,962 – 6,379,077 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 117 |
| Reading direction | reverse |
| Mean pairwise identity | 83.46 |
| Mean single sequence MFE | -29.68 |
| Consensus MFE | -19.25 |
| Energy contribution | -20.63 |
| Covariance contribution | 1.38 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.65 |
| SVM decision value | 0.38 |
| SVM RNA-class probability | 0.711592 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6378962 115 - 20766785 UA--GACAUGUGUAUUGACUAUAGAUCAUUGCUCGUUUUCGAUGAACAUGAACUUGAUGGGUUCAUAAAUAGCUUACAGACAGUUCUUGGGCAUGGAUACAGAUGCCCCACACAUGU ..--.((((((((.....((((.(.((((((........)))))).)(((((((.....)))))))..))))................((((((........)))))).)))))))) ( -30.50) >DroSec_CAF1 19625 115 - 1 UA--GACAUGUCUAUUGUCUAUAGACCAUUACUCGUUUGCGAUGAACAUGAACUUGACGGGUUCAUAAAUAGCUUACAGACAGUUCUUGGGCAUGGAUACAGAUGCCCCACACAUGU ..--.((((((..(((((((.(((..(((..(......)..)))...(((((((.....))))))).......))).)))))))....((((((........))))))...)))))) ( -29.00) >DroEre_CAF1 20836 102 - 1 CG--GGCAUGU-----GUCUAUAGAUCAUUGUUUGGUUUCGAUGAACAUGAACUUGAUGAGUUCAUAAAUAGCUUA----CACUUCUUGGG----GAUACACAUGCCCCACACAUGU .(--(((((((-----((((..(((((((((........))))))..((((((((...))))))))..........----....)))...)----))..)))))))))......... ( -31.40) >DroYak_CAF1 20898 104 - 1 UAGAGACAUGU-----GCCUAUAGAUCAUUGCUCGGUUUCGAUGAAUAUGAACUUGAUGAGUUCAUAAAUAGCUGA----CACUUCUUGGG----GAUACACAUGCCCCACACAUGU .....((((((-----(.((((...((((((........)))))).(((((((((...))))))))).))))....----.......((((----(.........)))))))))))) ( -27.80) >consensus UA__GACAUGU_____GUCUAUAGAUCAUUGCUCGGUUUCGAUGAACAUGAACUUGAUGAGUUCAUAAAUAGCUUA____CACUUCUUGGG____GAUACACAUGCCCCACACAUGU .....((((((.....(.((((.(.((((((........)))))).)(((((((.....)))))))..)))))...............((((((........))))))...)))))) (-19.25 = -20.63 + 1.38)

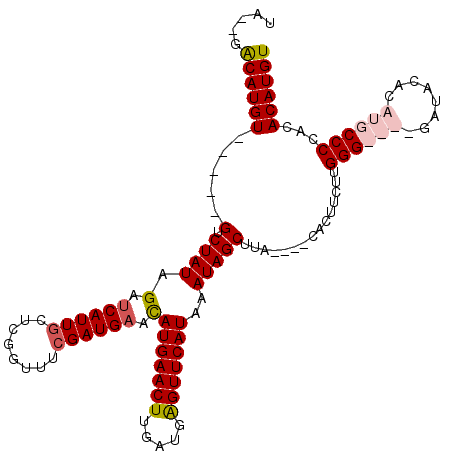

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:36 2006