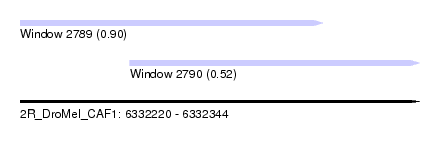
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,332,220 – 6,332,344 |
| Length | 124 |
| Max. P | 0.895632 |

| Location | 6,332,220 – 6,332,314 |
|---|---|
| Length | 94 |
| Sequences | 5 |
| Columns | 94 |
| Reading direction | forward |
| Mean pairwise identity | 92.13 |
| Mean single sequence MFE | -37.30 |
| Consensus MFE | -29.56 |
| Energy contribution | -30.76 |
| Covariance contribution | 1.20 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.93 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.99 |
| SVM RNA-class probability | 0.895632 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
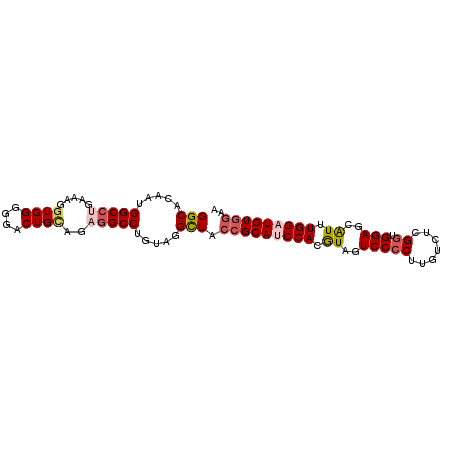
>2R_DroMel_CAF1 6332220 94 + 20766785 GGCACAAUGGCCUGAAAGGUGGGGGACUGCAGAGGCCUAUAGGCUACCGCAUCCACGUAGUCCACUUGUCUUGGUGGAGCAUUUGGAUGUGGAA (((.....(((((.....(..(....)..)..))))).....))).(((((((((.((..((((((......))))))..)).))))))))).. ( -42.90) >DroSec_CAF1 2233 94 + 1 GGCACAAUGGCCUGAAAGGUGGGGGACUGCAGAGGCCUGUAGGCUACCGCAUCCACAUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAA (((.....(((((.....(..(....)..)..))))).....))).(((((((((.((..(((((.......)).)))..)).))))))))).. ( -39.10) >DroSim_CAF1 2251 94 + 1 GGCACAAUGGCCUGAAAGGUGGGGGACUGCAGAGGCCUGUAGGCUACCGCAUCCACGUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAA (((.....(((((.....(..(....)..)..))))).....))).(((((((((.((..(((((.......)).)))..)).))))))))).. ( -39.40) >DroEre_CAF1 2269 94 + 1 GGCACAAUGGCGUGAAAGGUGGGGGACUGUAGUGGCCUGUAGGCUGCGGCAUCCACGUAGUCCCCUGGUCUCGGAGGAGCAUUUGGUUGUGAAA ..((((((.(.(((......(((((((((..(..(((....)))..)((...))...)))))))))...(((....)))))).).))))))... ( -34.40) >DroYak_CAF1 2283 94 + 1 GGCACAAUGGCCUGAAAGGUGGGGGACUGAAGCGGCCUGUAGGUUACCGCAUCCACGUAGUCCCCUUGUCUCGGAGGAGCGUUUGGUUGUGGUA ..((((((.(((.....)))((((((((((.((((((....))...)))).)).....)))))))).(.(((....)))).....))))))... ( -30.70) >consensus GGCACAAUGGCCUGAAAGGUGGGGGACUGCAGAGGCCUGUAGGCUACCGCAUCCACGUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAA (((.....(((((.....((((....))))..))))).....))).(((((((((.((..(((((.......)).)))..)).))))))))).. (-29.56 = -30.76 + 1.20)


| Location | 6,332,254 – 6,332,344 |
|---|---|
| Length | 90 |
| Sequences | 5 |
| Columns | 90 |
| Reading direction | forward |
| Mean pairwise identity | 92.89 |
| Mean single sequence MFE | -28.30 |
| Consensus MFE | -22.10 |
| Energy contribution | -22.86 |
| Covariance contribution | 0.76 |
| Combinations/Pair | 1.14 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.78 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.520354 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
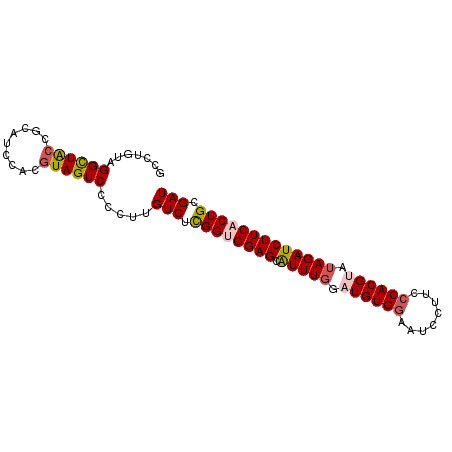
>2R_DroMel_CAF1 6332254 90 + 20766785 GCCUAUAGGCUACCGCAUCCACGUAGUCCACUUGUCUUGGUGGAGCAUUUGGAUGUGGAAUCCUUCCCACGUAAAGAUCUUCACUGCGAU (((....)))..(((((((((.((..((((((......))))))..)).)))))))))...........((((..(.....)..)))).. ( -31.00) >DroSec_CAF1 2267 90 + 1 GCCUGUAGGCUACCGCAUCCACAUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAAUCCUUCCCACGUAUAGAUCUUCACUGCGAU (((....))).......................(((.((((((((.(((((.((((((........)))))).))))))))))))).))) ( -27.40) >DroSim_CAF1 2285 90 + 1 GCCUGUAGGCUACCGCAUCCACGUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAAUCCUUCCCACGUAUAGAUCUUCACUGCGAU ....(..((((((.........))))))..)..(((.((((((((.(((((.((((((........)))))).))))))))))))).))) ( -28.70) >DroEre_CAF1 2303 90 + 1 GCCUGUAGGCUGCGGCAUCCACGUAGUCCCCUGGUCUCGGAGGAGCAUUUGGUUGUGAAAUCCUUCCCACGUAUAGAUCUUCACUGCGAU (((.(..(((((((.......)))))))..).))).(((.(((((.(((((..((((..........))))..))))).))).)).))). ( -24.90) >DroYak_CAF1 2317 90 + 1 GCCUGUAGGUUACCGCAUCCACGUAGUCCCCUUGUCUCGGAGGAGCGUUUGGUUGUGGUAUCCUACCCACGUAUAGAUCUUCACUGCGAU ....(((((.(((((((.((((((..(((((.......)).)))))))..)).))))))).)))))...((((..(.....)..)))).. ( -29.50) >consensus GCCUGUAGGCUACCGCAUCCACGUAGUCCCCUUGUCUCGGUGGAGCAUUUGGAUGUGGAAUCCUUCCCACGUAUAGAUCUUCACUGCGAU .......((((((.........)))))).....(((.((((((((.(((((.((((((........)))))).))))))))))))).))) (-22.10 = -22.86 + 0.76)

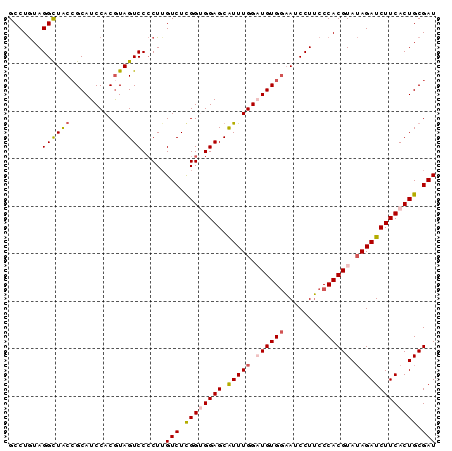
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:10:17 2006