
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 6,114,792 – 6,114,892 |
| Length | 100 |
| Max. P | 0.999953 |

| Location | 6,114,792 – 6,114,892 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | forward |
| Mean pairwise identity | 93.54 |
| Mean single sequence MFE | -30.64 |
| Consensus MFE | -25.94 |
| Energy contribution | -26.02 |
| Covariance contribution | 0.08 |
| Combinations/Pair | 1.08 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.85 |
| SVM decision value | 0.83 |
| SVM RNA-class probability | 0.860280 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6114792 100 + 20766785 CGUUUUGUCUCCCACACCGUGAGAUGCAAAAGCACAAAGUGCAUAUGCUAGCAGCUGC--CGGUGUCCGCUUAACAGCUGCUUAACAUGCUUACUGCUGCUG ...(((((.((.(((...))).)).)))))((((...((((..((((..((((((((.--.(((....)))...))))))))...))))..))))..)))). ( -30.90) >DroSec_CAF1 123883 97 + 1 CGUUUUGUCUCCCACACCGAGAGAUGCUAAAGCACAAAGUGCAUAUGCUAGCAGCUGC--CGGUGUCCGCUUAACAGCUGCUUAACGUGCUUACUG---CUG ......(((((.(.....).)))))((..((((((.....((....)).((((((((.--.(((....)))...))))))))....))))))...)---).. ( -29.80) >DroSim_CAF1 129973 97 + 1 CGUUUUGUCUCCCACACCGAGAGAUGCUAAAGCACAAAGUGCAUAUGCUAGCAGCUGC--CGGUGUCCGCUUAACAGCUGCUUAACGUGCUUACUG---CUG ......(((((.(.....).)))))((..((((((.....((....)).((((((((.--.(((....)))...))))))))....))))))...)---).. ( -29.80) >DroEre_CAF1 127551 99 + 1 CGUUUUGUCUCCCACACCGAAAGAUGCAAAAGCACAAAGUGCAUAUGCUAGCAGCUGCCCCGGUGUCCACUUAACAGCUGCUUAACGUGCUUAGUG---CUG ...(((((.((...........)).)))))(((((.(((..(.......((((((((....(((....)))...))))))))....)..))).)))---)). ( -28.80) >DroYak_CAF1 125935 97 + 1 CGUCUUGUCUCCCACACCGAGAGAUGCAGGAGCACAAAGUGCAUUUGCUAGCAGCUGC--CGGUGUCCGCUUAACAGCUGCUUAACGUGCUUAGUG---CUG ..(((((((((.(.....).)))..))))))((((.(((..(.((....((((((((.--.(((....)))...)))))))).)).)..))).)))---).. ( -33.90) >consensus CGUUUUGUCUCCCACACCGAGAGAUGCAAAAGCACAAAGUGCAUAUGCUAGCAGCUGC__CGGUGUCCGCUUAACAGCUGCUUAACGUGCUUACUG___CUG ......(((((.(.....).)))))....((((((.....((....)).((((((((....(((....)))...))))))))....)))))).......... (-25.94 = -26.02 + 0.08)


| Location | 6,114,792 – 6,114,892 |
|---|---|
| Length | 100 |
| Sequences | 5 |
| Columns | 102 |
| Reading direction | reverse |
| Mean pairwise identity | 93.54 |
| Mean single sequence MFE | -36.18 |
| Consensus MFE | -34.64 |
| Energy contribution | -34.00 |
| Covariance contribution | -0.64 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.55 |
| Structure conservation index | 0.96 |
| SVM decision value | 4.82 |
| SVM RNA-class probability | 0.999953 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 6114792 100 - 20766785 CAGCAGCAGUAAGCAUGUUAAGCAGCUGUUAAGCGGACACCG--GCAGCUGCUAGCAUAUGCACUUUGUGCUUUUGCAUCUCACGGUGUGGGAGACAAAACG ..(((((.....))((((((.((((((((....(((...)))--)))))))))))))).)))..(((((.((((..((((....))))..)))))))))... ( -38.30) >DroSec_CAF1 123883 97 - 1 CAG---CAGUAAGCACGUUAAGCAGCUGUUAAGCGGACACCG--GCAGCUGCUAGCAUAUGCACUUUGUGCUUUAGCAUCUCUCGGUGUGGGAGACAAAACG ..(---(...(((((((...(((((((((....(((...)))--))))))))).((....))....)))))))..)).((((((.....))))))....... ( -34.80) >DroSim_CAF1 129973 97 - 1 CAG---CAGUAAGCACGUUAAGCAGCUGUUAAGCGGACACCG--GCAGCUGCUAGCAUAUGCACUUUGUGCUUUAGCAUCUCUCGGUGUGGGAGACAAAACG ..(---(...(((((((...(((((((((....(((...)))--))))))))).((....))....)))))))..)).((((((.....))))))....... ( -34.80) >DroEre_CAF1 127551 99 - 1 CAG---CACUAAGCACGUUAAGCAGCUGUUAAGUGGACACCGGGGCAGCUGCUAGCAUAUGCACUUUGUGCUUUUGCAUCUUUCGGUGUGGGAGACAAAACG ...---......(((.((((.(((((((((..(.(....))..)))))))))))))...)))..(((((.((((..((((....))))..)))))))))... ( -34.90) >DroYak_CAF1 125935 97 - 1 CAG---CACUAAGCACGUUAAGCAGCUGUUAAGCGGACACCG--GCAGCUGCUAGCAAAUGCACUUUGUGCUCCUGCAUCUCUCGGUGUGGGAGACAAGACG ...---......(((.((((.((((((((....(((...)))--))))))))))))...)))..(((((.((((..((((....))))..)))))))))... ( -38.10) >consensus CAG___CAGUAAGCACGUUAAGCAGCUGUUAAGCGGACACCG__GCAGCUGCUAGCAUAUGCACUUUGUGCUUUUGCAUCUCUCGGUGUGGGAGACAAAACG ............(((.((((.((((((((....(((...)))..))))))))))))...)))..(((((.((((..((((....))))..)))))))))... (-34.64 = -34.00 + -0.64)

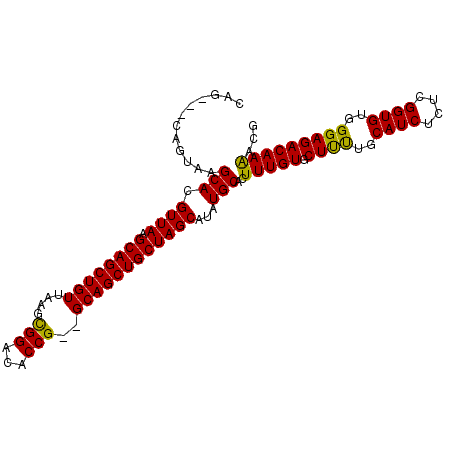

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:09:15 2006