
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,933,158 – 5,933,248 |
| Length | 90 |
| Max. P | 0.744307 |

| Location | 5,933,158 – 5,933,248 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | forward |
| Mean pairwise identity | 61.78 |
| Mean single sequence MFE | -18.23 |
| Consensus MFE | -8.25 |
| Energy contribution | -9.50 |
| Covariance contribution | 1.25 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744307 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5933158 90 + 20766785 UUU--------------CAAAUAAGCCAG--AAUUGGUGGCUAAUUUCCAAAGUUGUGCAUGCAAUCCAAACAGCUGAUCCAA--CAGCUGAUCAUUUUCCAAAGAGU .((--------------(((((.(((((.--......))))).)))).....(((((....))))).....((((((......--)))))).............))). ( -17.50) >DroSec_CAF1 75617 90 + 1 UUU--------------CAAAUAAGCCAG--AAUUGCUGGCUAAUUUCAAAAGUUGUCCAUGUAAACCAAACAGCUGACCCAA--CAGCUGAUCACUUUCCAAAGAGU (((--------------.((((.((((((--.....)))))).)))).)))....................((((((......--))))))...((((......)))) ( -20.00) >DroSim_CAF1 74164 100 + 1 UUUGCUUUAAACUUGAUCAAAUAACCCAAUCAGUCCCGAACCAAUUCCUUUAGG---AUCUGGUAUCAAAACAGCUGAUCCGAUUCAGCCGG-----CUUCAAACAUA ...............................((((..((((((..(((....))---)..)))).))......(((((......))))).))-----))......... ( -18.60) >DroYak_CAF1 78280 83 + 1 UU---------------CCAGUAAGCCUG--AAUUAGUGGCUAUUUUGAACAGCUUCACAU------CAAACAGCUGAUCCAU--CAGCUGAUUACUUUCCAAAGAGU ..---------------..((((((((..--.......))))...................------....(((((((....)--))))))..))))........... ( -16.80) >consensus UUU______________CAAAUAAGCCAG__AAUUGCUGGCUAAUUUCAAAAGUUGUACAUG_AAUCCAAACAGCUGAUCCAA__CAGCUGAUCACUUUCCAAAGAGU .......................((((...........)))).............................((((((........))))))................. ( -8.25 = -9.50 + 1.25)


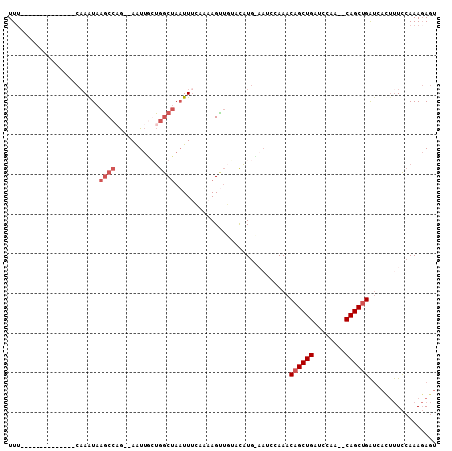
| Location | 5,933,158 – 5,933,248 |
|---|---|
| Length | 90 |
| Sequences | 4 |
| Columns | 108 |
| Reading direction | reverse |
| Mean pairwise identity | 61.78 |
| Mean single sequence MFE | -23.90 |
| Consensus MFE | -9.89 |
| Energy contribution | -10.95 |
| Covariance contribution | 1.06 |
| Combinations/Pair | 1.38 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.41 |
| SVM decision value | -0.02 |
| SVM RNA-class probability | 0.522106 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5933158 90 - 20766785 ACUCUUUGGAAAAUGAUCAGCUG--UUGGAUCAGCUGUUUGGAUUGCAUGCACAACUUUGGAAAUUAGCCACCAAUU--CUGGCUUAUUUG--------------AAA ......((.((..(((.((((((--......)))))).)))..)).))........((..((....(((((......--.)))))..))..--------------)). ( -17.70) >DroSec_CAF1 75617 90 - 1 ACUCUUUGGAAAGUGAUCAGCUG--UUGGGUCAGCUGUUUGGUUUACAUGGACAACUUUUGAAAUUAGCCAGCAAUU--CUGGCUUAUUUG--------------AAA ..(((.((.(((.(((.((((((--......)))))).))).))).)).))).....(((.((((.((((((.....--)))))).)))).--------------))) ( -24.90) >DroSim_CAF1 74164 100 - 1 UAUGUUUGAAG-----CCGGCUGAAUCGGAUCAGCUGUUUUGAUACCAGAU---CCUAAAGGAAUUGGUUCGGGACUGAUUGGGUUAUUUGAUCAAGUUUAAAGCAAA ..(((((.(((-----(.((((((......))))))(((((((.(((((.(---((....))).))))))))))))((((..(.....)..)))).)))).))))).. ( -30.70) >DroYak_CAF1 78280 83 - 1 ACUCUUUGGAAAGUAAUCAGCUG--AUGGAUCAGCUGUUUG------AUGUGAAGCUGUUCAAAAUAGCCACUAAUU--CAGGCUUACUGG---------------AA ....((..(.((((...((((((--(....))))))).(((------(.(((..(((((.....))))))))....)--))))))).)..)---------------). ( -22.30) >consensus ACUCUUUGGAAAGUGAUCAGCUG__UUGGAUCAGCUGUUUGGAUU_CAUGUACAACUUUUGAAAUUAGCCACCAAUU__CUGGCUUAUUUG______________AAA .................((((((........))))))........................((((.((((((.......)))))).)))).................. ( -9.89 = -10.95 + 1.06)

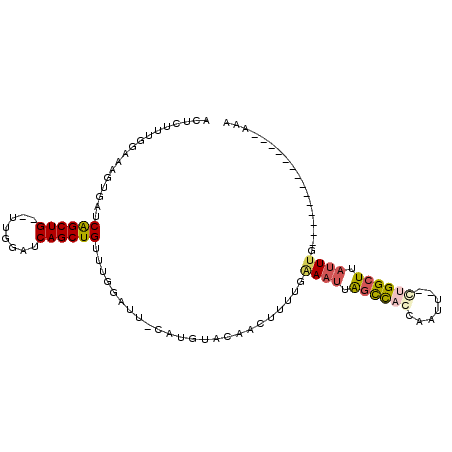
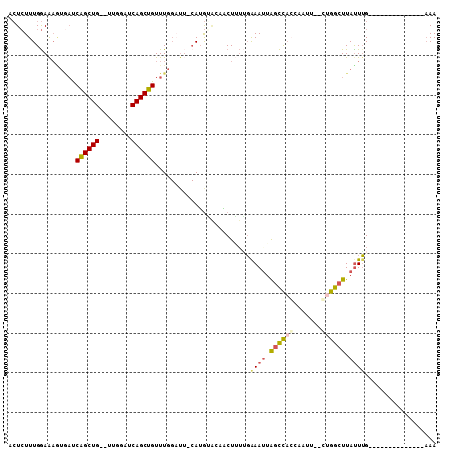
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:07:54 2006