
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,811,343 – 5,811,457 |
| Length | 114 |
| Max. P | 0.744578 |

| Location | 5,811,343 – 5,811,457 |
|---|---|
| Length | 114 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.22 |
| Mean single sequence MFE | -44.37 |
| Consensus MFE | -34.04 |
| Energy contribution | -32.82 |
| Covariance contribution | -1.22 |
| Combinations/Pair | 1.19 |
| Mean z-score | -1.99 |
| Structure conservation index | 0.77 |
| SVM decision value | 0.46 |
| SVM RNA-class probability | 0.744578 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
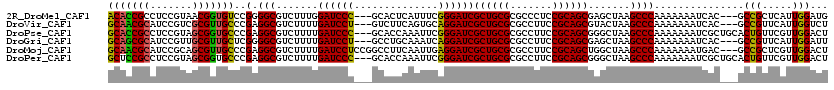
>2R_DroMel_CAF1 5811343 114 - 20766785 ACACCGCCUCCGUAACGGUGUCCGGGGCGUCUUUGGAUCCC---GCACUCAUUUCGGGAUCGCUGCGCGCCCUCCGCAGCGAGCUAAGCCCAAAAAAAUCAC---GCCGCUCAUUGGAUG ........((((...((((((...((((.......((((((---(.........)))))))((((((.......)))))).......)))).........))---)))).....)))).. ( -38.70) >DroVir_CAF1 7501 114 - 1 GCAACGCAUCCGUCGCGUUGCCCGAGGCGUCUUUUGAUCCU---GUCUUCAGUGCAGGAUCGCUGCGCGCCUUCCGCAGCGUACUAAGCCCAAAAAAAUCAC---GCCGUUCAUUGGUCU (((((((.......)))))))(((((((((..(((((((((---((.......))))))))((((((.......))))))...............)))..))---))).....))))... ( -42.40) >DroPse_CAF1 7414 117 - 1 GCACCGCCUCCGUAGCGGUGCCCGAGGCGUCUUUUGAUCCC---GCACCAAAUUCGGGAUCGCUGCGCGCCUUCCGCAGCGGGCUAAGCCCAAAAAAAUCGCUGCACUGUUCGUUGGACU (((((((.......)))))))(((((((((.(..(((((((---(.........))))))))..).)))))))..(((((((((...)))).........)))))..........))... ( -49.00) >DroGri_CAF1 8126 114 - 1 GCAGCGCAUCCGUUGCGUUGCUCGGGGCGUCUUUUGAUCCU---GCCUGCAAAUCAGGAUCGCUGCGCGCCUUCCGCAGCGAGCUAAGCCCAAAAAAAUCAC---GCCGUUCAUUGGAUU ((((((((.....))))))))...((((.......((((((---(.........)))))))((((((.......)))))).......))))...........---.(((.....)))... ( -43.10) >DroMoj_CAF1 17335 117 - 1 GCAACGCAUCCGCAGCGUUGCCCGAGGCGUCUUUUGAUCCUCCGGCCUUCAAUUGAGGAUCGCUGCGCGCCUUCCGCAGCUGGCUAAGCCCAAAAAAAUGAC---GCCGCUCGUUGGACU (((((((.......)))))))(((((((((((((((((((((.(........).)))))))((((((.......)))))).(((...)))....)))).)))---))).....))))... ( -45.00) >DroPer_CAF1 7455 117 - 1 GCUCCGCCUCCGUAGCGGUGCCCGAGGCGUCUUUUGAUCCC---GCACCAAAUUCGGGAUCGCUGCGCGCCUUCCGCAGCGGGCUAAGCCCAAAAAAAUCGCUGCACUGUUCGUUGGACU ......((..((.((((((((....((((..((((((((((---(.........)))))))((((((.......))))))((((...))))..))))..))))))))))))))..))... ( -48.00) >consensus GCACCGCAUCCGUAGCGGUGCCCGAGGCGUCUUUUGAUCCC___GCACUCAAUUCAGGAUCGCUGCGCGCCUUCCGCAGCGGGCUAAGCCCAAAAAAAUCAC___GCCGUUCAUUGGACU (((((((.......)))))))..(.(((.......((((((..............))))))((((((.......)))))).......))))...............(((.....)))... (-34.04 = -32.82 + -1.22)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:07:03 2006