
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,665,088 – 5,665,200 |
| Length | 112 |
| Max. P | 0.751675 |

| Location | 5,665,088 – 5,665,200 |
|---|---|
| Length | 112 |
| Sequences | 6 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 76.40 |
| Mean single sequence MFE | -31.73 |
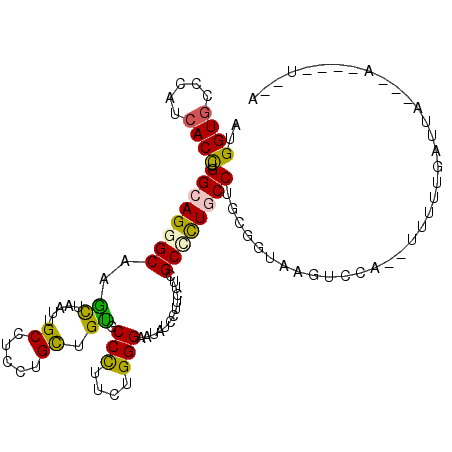
| Consensus MFE | -19.63 |
| Energy contribution | -18.97 |
| Covariance contribution | -0.66 |
| Combinations/Pair | 1.42 |
| Mean z-score | -1.51 |
| Structure conservation index | 0.62 |
| SVM decision value | 0.48 |
| SVM RNA-class probability | 0.751675 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
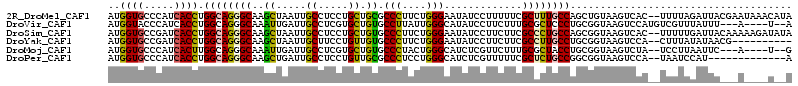
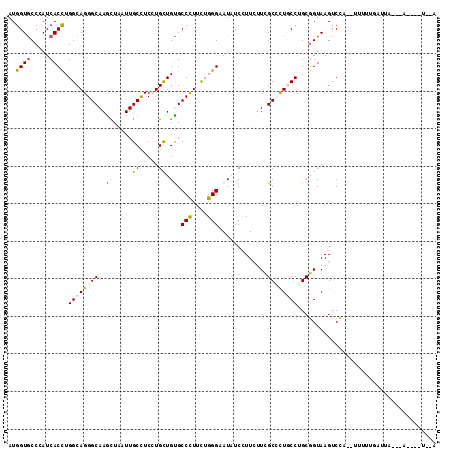
>2R_DroMel_CAF1 5665088 112 - 20766785 AUGGUGCCCAUCACCUGGCAGGGCAAGCUAAUUGCCUCCUGCUGCGCCCUUCUGGGAAUAUCCUUUUUCGCUUUGCCAGCUGUAAGUCAC--UUUUAGAUUACGAAUAAACAUA ..((((.(...((.((((((((((.(((............)))...(((....))).............)))))))))).))...).)))--)..................... ( -28.10) >DroVir_CAF1 2949 105 - 1 AUGGUACCCAUCACCUGGCAGGGCAAAUUGAUUGCCUCGUGCUGUGCCUUAUUGGGCAUAUCCUUCUUUGCGCUCCCUGCGGUAAGUCCAUGUCGUUUAUUU---A----U--A ((((((((((.....))(((((((((.....)))))..(((((((((((....))))))).........))))...))))))))...))))...........---.----.--. ( -28.60) >DroSim_CAF1 2494 112 - 1 AUGGUGCCGAUCACCUGGCAGGGCAAGCUAAUUGCCUCCUGCUGUGCCCUUCUGGGAAUAUCCUUCUUCGCCCUGCCAGCGGUAAGUCAC--UUUUUGAUUACAAAAAGAUAUA ....(((((.....((((((((((.(((............)))...(((....))).............))))))))))))))).....(--((((((....)))))))..... ( -38.70) >DroYak_CAF1 2360 102 - 1 AUGGUGCCGAUCACCUGGCAGGGCAAGCUAAUUGCUUCCUGUUGUGCCCUUCUGGGAAUAUCCUUCUUCGCCUUGCCUGCGGUAAGUCCA--CUUUAUAUAACG---------- .((((((((.......((((((((((((.....)))).....(((.(((....))).))).........))))))))..)))))...)))--............---------- ( -31.00) >DroMoj_CAF1 2914 103 - 1 AUGGUGCCCAUCACUUGGCAGGGCAAAUUGAUUGCCUCGUGCUGUGCCCUACUGGGCAUCUCGUUCUUUGCGCUACCUGCGGUAAGUCUA--UCCUUAAUUC---A----U--G ..((((((((......((((.((((...(((.....))))))).))))....))))))))..........(((.....))).((((....--..))))....---.----.--. ( -29.30) >DroPer_CAF1 2310 99 - 1 AUGGUGCCCAUCACCUGGCAGGGCAAGCUGAUUGCCUCCUGUUGCGCCCUCCUGGGCAUCUCGUUUUUCGCUCUGCCGGCGGUAAGUCCA--UAAUCCAU-------------A ((((.((..(((.((.((((((((..((.....))..........((((....))))............)))))))))).)))..)))))--).......-------------. ( -34.70) >consensus AUGGUGCCCAUCACCUGGCAGGGCAAGCUAAUUGCCUCCUGCUGUGCCCUUCUGGGAAUAUCCUUCUUCGCCCUGCCUGCGGUAAGUCCA__UUUUUGAUUA___A____U__A ..((((.....)))).((((((((..((.....((.....)).)).(((....))).............))))))))..................................... (-19.63 = -18.97 + -0.66)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:06:07 2006