
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,516,535 – 5,516,627 |
| Length | 92 |
| Max. P | 0.999788 |

| Location | 5,516,535 – 5,516,627 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | forward |
| Mean pairwise identity | 90.44 |
| Mean single sequence MFE | -31.21 |
| Consensus MFE | -24.32 |
| Energy contribution | -24.36 |
| Covariance contribution | 0.04 |
| Combinations/Pair | 1.13 |
| Mean z-score | -4.49 |
| Structure conservation index | 0.78 |
| SVM decision value | 3.01 |
| SVM RNA-class probability | 0.998116 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5516535 92 + 20766785 GCCAAAUAAAUCUUGAUUUCCCCCGACCUAUUCAUGGCCAAAAAAAUUGGCCAGAAUUGUCGGGGGAAUUACACCAGCAGAUUGUCUAUAUG--- ........(((((((..((((((((((..((((.(((((((.....))))))))))).))))))))))...)).....))))).........--- ( -35.70) >DroSec_CAF1 40986 92 + 1 GCCAAAUAAAUCUUGAUUUCCCCCGACCUAUUCACGGCCAAAAAAAUUGGCAGGAAUUGUUGGGGGAAUUACACCAGCAGAUUGUCUAUAUG--- ........(((((((..((((((((((..((((...(((((.....)))))..)))).))))))))))...)).....))))).........--- ( -29.80) >DroSim_CAF1 18196 92 + 1 GCCAAAUAAAUCUUGAUUUCCCCCGACCUAUUCACGGCCAAAAAAAUUGGCCAGAAUUGUUGGGGGAAUUACACCAGCAGAUUGUCUAUAUG--- ........(((((((..((((((((((..((((..((((((.....)))))).)))).))))))))))...)).....))))).........--- ( -31.00) >DroEre_CAF1 17847 94 + 1 ACCGAAUAAAUCUUGAUUUCC-CCGACCUAUUCACGGCCAAAAAAGUUGGCCAAAAAUGUCGGGGGAAUUACACCAGCAGAUUGUCCAUAUACAU ...((...(((((((((((((-(((((........((((((.....))))))......))))))))))))).......))))).))......... ( -30.35) >DroYak_CAF1 18404 90 + 1 GCCGAAUAAAUCUUGAUUUCC-CCGAUCUAUUCACGGCCAAAAAA-UUGGCCUCAAAUGUCGGGGGGAUUACACCAGCAGAUUGUCUAUUUG--- ..(((((((((((((((..((-(((((..(((...((((((....-))))))...))))))))))..)))).......)))))...))))))--- ( -29.21) >consensus GCCAAAUAAAUCUUGAUUUCCCCCGACCUAUUCACGGCCAAAAAAAUUGGCCAGAAUUGUCGGGGGAAUUACACCAGCAGAUUGUCUAUAUG___ ........(((((((..((((((((((........((((((.....))))))......))))))))))...)).....)))))............ (-24.32 = -24.36 + 0.04)


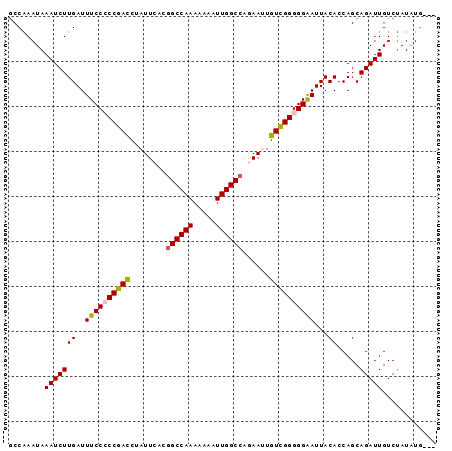
| Location | 5,516,535 – 5,516,627 |
|---|---|
| Length | 92 |
| Sequences | 5 |
| Columns | 95 |
| Reading direction | reverse |
| Mean pairwise identity | 90.44 |
| Mean single sequence MFE | -30.30 |
| Consensus MFE | -26.20 |
| Energy contribution | -27.64 |
| Covariance contribution | 1.44 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.20 |
| Structure conservation index | 0.86 |
| SVM decision value | 4.08 |
| SVM RNA-class probability | 0.999788 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5516535 92 - 20766785 ---CAUAUAGACAAUCUGCUGGUGUAAUUCCCCCGACAAUUCUGGCCAAUUUUUUUGGCCAUGAAUAGGUCGGGGGAAAUCAAGAUUUAUUUGGC ---((.(((...(((((..((((....((((((((((.(((((((((((.....))))))).))))..)))))))))))))))))))))).)).. ( -37.60) >DroSec_CAF1 40986 92 - 1 ---CAUAUAGACAAUCUGCUGGUGUAAUUCCCCCAACAAUUCCUGCCAAUUUUUUUGGCCGUGAAUAGGUCGGGGGAAAUCAAGAUUUAUUUGGC ---((.(((...(((((..((((....(((((((.((.(((((.(((((.....))))).).))))..)).))))))))))))))))))).)).. ( -27.10) >DroSim_CAF1 18196 92 - 1 ---CAUAUAGACAAUCUGCUGGUGUAAUUCCCCCAACAAUUCUGGCCAAUUUUUUUGGCCGUGAAUAGGUCGGGGGAAAUCAAGAUUUAUUUGGC ---((.(((...(((((..((((....(((((((.((.(((((((((((.....))))))).))))..)).))))))))))))))))))).)).. ( -31.00) >DroEre_CAF1 17847 94 - 1 AUGUAUAUGGACAAUCUGCUGGUGUAAUUCCCCCGACAUUUUUGGCCAACUUUUUUGGCCGUGAAUAGGUCGG-GGAAAUCAAGAUUUAUUCGGU .(((......)))....(((((.((((.(((((((((((((.(((((((.....))))))).))))..)))))-)).......)).))))))))) ( -28.41) >DroYak_CAF1 18404 90 - 1 ---CAAAUAGACAAUCUGCUGGUGUAAUCCCCCCGACAUUUGAGGCCAA-UUUUUUGGCCGUGAAUAGAUCGG-GGAAAUCAAGAUUUAUUCGGC ---..............(((((...((((.(((((((((((..((((((-....))))))..)))).).))))-)).......))))...))))) ( -27.40) >consensus ___CAUAUAGACAAUCUGCUGGUGUAAUUCCCCCGACAAUUCUGGCCAAUUUUUUUGGCCGUGAAUAGGUCGGGGGAAAUCAAGAUUUAUUUGGC ............(((((..((((....((((((((((..((((((((((.....))))))).)))...)))))))))))))))))))........ (-26.20 = -27.64 + 1.44)
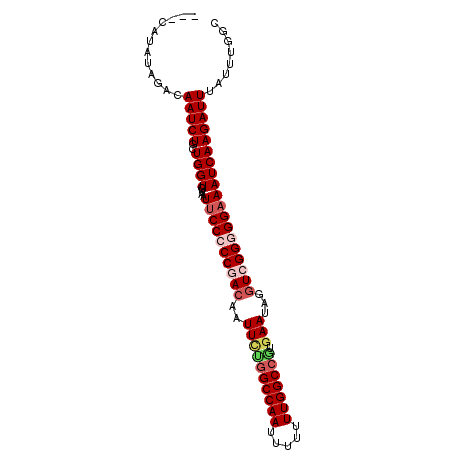



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:53 2006