
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,435,731 – 5,435,850 |
| Length | 119 |
| Max. P | 0.999710 |

| Location | 5,435,731 – 5,435,850 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | forward |
| Mean pairwise identity | 93.77 |
| Mean single sequence MFE | -37.11 |
| Consensus MFE | -33.78 |
| Energy contribution | -34.02 |
| Covariance contribution | 0.24 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.36 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880810 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5435731 119 + 20766785 CUUAGGACGUGCCACUCACCGUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAAUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGU ......(((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..))) ( -36.72) >DroSec_CAF1 24388 119 + 1 CUUAGGACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGU ......(((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..))) ( -36.72) >DroSim_CAF1 24522 119 + 1 CUUAGGACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGACUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGU ......(((.((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..))) ( -36.72) >DroEre_CAF1 24840 117 + 1 CUUAGGAUGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAGCUCGGAAUAGGCUUGCGAUGGA--GCAACCGGCAGUAAUUCAUGUUUUAGUCACCAUUUGAGUGGCAUUAU ........((((((((((.......(((((((((((((.(((......))).)))).((.((((......--)))))))))).(((........))))))))....))))))))))... ( -37.80) >DroYak_CAF1 25276 117 + 1 CUUAGGAUGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGA--GCCCCCGGCAGUAACUCAUGUUUAAGUCACUUCUUGAGUGGCUUUGU ..........((((((((.......(((((((((((((..(((.....))).)))).(((((......))--)))...)))...(((....)))..))))))....))))))))..... ( -37.60) >consensus CUUAGGACGUGCCACUCACCAUUUUGUGACUGCCGUUCAGGGGAUUAUCUCGGAAUAGGCUUGCGAUUGAACGCCCCCGGCAGUAACUCAUGUUUUAGUCACUUCUUGAGUGGCUUUGU ..........((((((((.......((((((((((....((((......(((.((.....)).))).......))))))))...(((....)))..))))))....))))))))..... (-33.78 = -34.02 + 0.24)

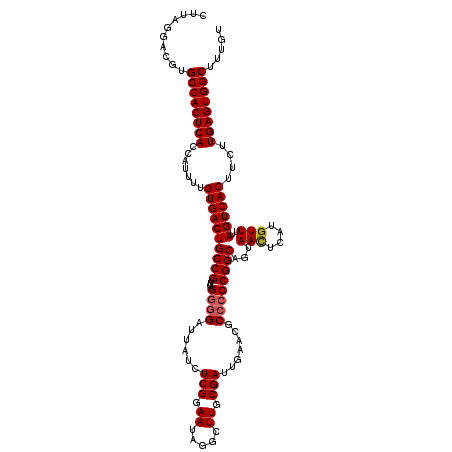
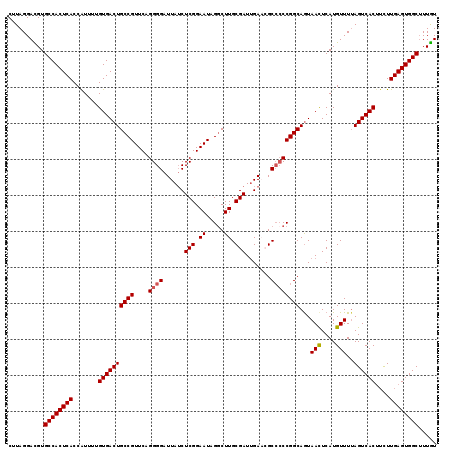
| Location | 5,435,731 – 5,435,850 |
|---|---|
| Length | 119 |
| Sequences | 5 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 93.77 |
| Mean single sequence MFE | -37.14 |
| Consensus MFE | -33.26 |
| Energy contribution | -33.86 |
| Covariance contribution | 0.60 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.59 |
| Structure conservation index | 0.90 |
| SVM decision value | 3.93 |
| SVM RNA-class probability | 0.999710 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5435731 119 - 20766785 ACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUAUUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAACGGUGAGUGGCACGUCCUAAG .....((((((((....((((((..(((....)))...((((((((.(((...(((.((.....)).)))...)))))))....)))))))))).......)))))))).......... ( -37.10) >DroSec_CAF1 24388 119 - 1 ACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGUCCUAAG .....((((((((....((((((..(((....)))...((((((((.(((...(((.((.....)).)))...)))))))....)))))))))).......)))))))).......... ( -37.10) >DroSim_CAF1 24522 119 - 1 ACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGUCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGUCCUAAG .....((((((((....((((((..(((....)))...((((((((((......)))..((((......))))...)))).....))))))))).......)))))))).......... ( -38.50) >DroEre_CAF1 24840 117 - 1 AUAAUGCCACUCAAAUGGUGACUAAAACAUGAAUUACUGCCGGUUGC--UCCAUCGCAAGCCUAUUCCGAGCUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACAUCCUAAG ....(((((((((...(....).....(((.....(((((((..(((--......)))((((......).)))...........)))))))......))).)))))))))......... ( -34.70) >DroYak_CAF1 25276 117 - 1 ACAAAGCCACUCAAGAAGUGACUUAAACAUGAGUUACUGCCGGGGGC--UCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACAUCCUAAG .....((((((((...(((((((((....)))))))))((((((((.--....(((.((.....)).)))......))))....)))).............)))))))).......... ( -38.30) >consensus ACAAAGCCACUCAAGAAGUGACUAAAACAUGAGUUACUGCCGGGGGCGUUCAAUCGCAAGCCUAUUCCGAGAUAAUCCCCUGAACGGCAGUCACAAAAUGGUGAGUGGCACGUCCUAAG .....((((((((....((((((..(((....)))...((((((((.......(((.((.....)).)))......))))....)))))))))).......)))))))).......... (-33.26 = -33.86 + 0.60)

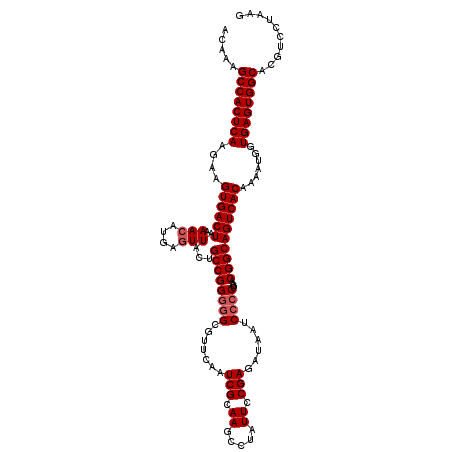
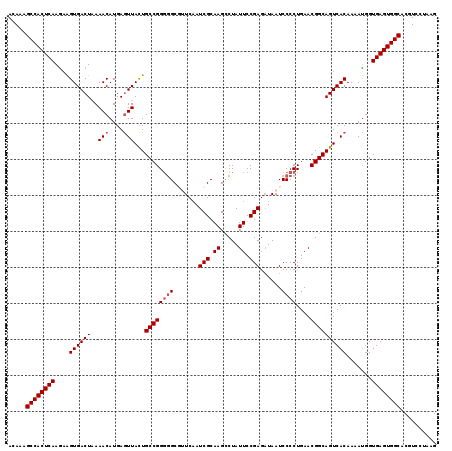
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:04:04 2006