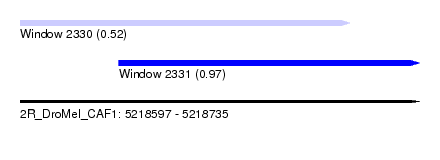
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,218,597 – 5,218,735 |
| Length | 138 |
| Max. P | 0.968854 |

| Location | 5,218,597 – 5,218,711 |
|---|---|
| Length | 114 |
| Sequences | 3 |
| Columns | 114 |
| Reading direction | forward |
| Mean pairwise identity | 91.81 |
| Mean single sequence MFE | -31.57 |
| Consensus MFE | -27.20 |
| Energy contribution | -27.43 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.15 |
| Mean z-score | -1.77 |
| Structure conservation index | 0.86 |
| SVM decision value | -0.03 |
| SVM RNA-class probability | 0.520808 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
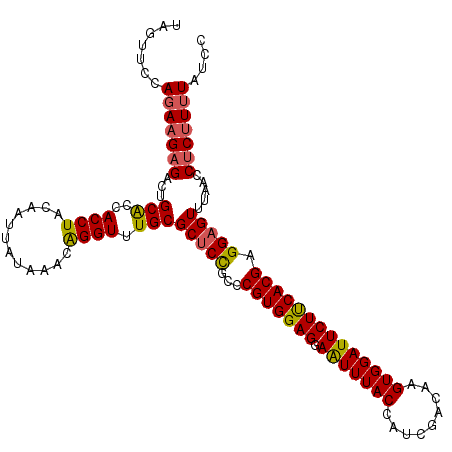
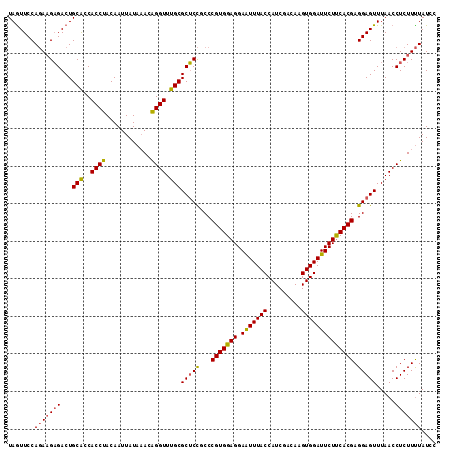
>2R_DroMel_CAF1 5218597 114 + 20766785 UAGUUCCACAAGUGACUGCGUCACCUACAAUUAUCAACGGGUUUGCGCACUGCCCGUGGAGGAGUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUCUAUCC ....(((((..(((((...)))))..............((((.........)))))))))((((..(.((((......)))))..))))..((((((......))))))..... ( -31.30) >DroSec_CAF1 94669 114 + 1 UAAUUCCAGAAGAGACUGCACCACCUACAAUUAUAUACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUCCACGAGGAGUUUAACCUCUUUUAUCC .......(((((((...(((..((((............)))).)))(((((...(((((((.(((((((.........)))))))))))))).))))).....))))))).... ( -34.70) >DroSim_CAF1 80478 114 + 1 UAGUUCCAGACGAGACUGCACCACCUACAAUUAUAAACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUUUAUCC ((((..(....)..))))..............(((((.(((((...(((((...((((.((((((((((.........)))))))))))))).)))))..)))))..))))).. ( -28.70) >consensus UAGUUCCAGAAGAGACUGCACCACCUACAAUUAUAAACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUUUAUCC .......(((((((...(((..((((............)))).)))(((((...(((((((.(((((((.........)))))))))))))).))))).....))))))).... (-27.20 = -27.43 + 0.23)

| Location | 5,218,631 – 5,218,735 |
|---|---|
| Length | 104 |
| Sequences | 3 |
| Columns | 104 |
| Reading direction | forward |
| Mean pairwise identity | 94.23 |
| Mean single sequence MFE | -34.90 |
| Consensus MFE | -32.47 |
| Energy contribution | -32.03 |
| Covariance contribution | -0.43 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.72 |
| Structure conservation index | 0.93 |
| SVM decision value | 1.63 |
| SVM RNA-class probability | 0.968854 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
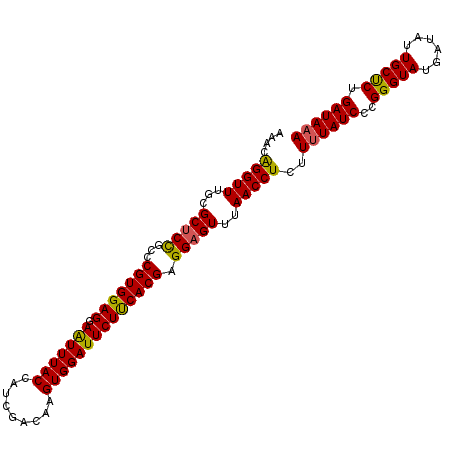
>2R_DroMel_CAF1 5218631 104 + 20766785 CAACGGGUUUGCGCACUGCCCGUGGAGGAGUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUCUAUCCCGGGUAUGAUAUUGCUCUGAUAAA ....((((.....((.((((((.(((((((..(.((((......)))))..))))..((((((......))))))..))))))))))).....))))....... ( -33.00) >DroSec_CAF1 94703 104 + 1 AUACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUCCACGAGGAGUUUAACCUCUUUUAUCCCGGGUAUGAUAUUGCCCUGAUAAA ....(((((...(((((...(((((((.(((((((.........)))))))))))))).)))))..)))))..((((((..(((((......))))).)))))) ( -38.70) >DroSim_CAF1 80512 104 + 1 AAACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUUUAUCCCGGGUAUGAUAUUGCUCUGAUAAA ....(((((...(((((...((((.((((((((((.........)))))))))))))).)))))..)))))..((((((..(((((......))))).)))))) ( -33.00) >consensus AAACAGGUUUGCGCUCCGCCCGUGGAGGAAUUUACCAUCGACAAGUGGAUUCUUCACGAGGAGUUUAACCUCUUUUAUCCCGGGUAUGAUAUUGCUCUGAUAAA ....(((((...(((((...(((((((.(((((((.........)))))))))))))).)))))..)))))..((((((..(((((......))))).)))))) (-32.47 = -32.03 + -0.43)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:02:48 2006