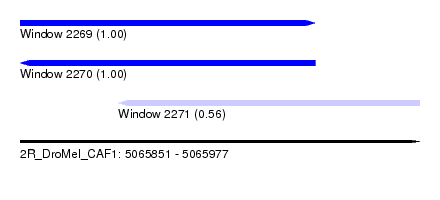

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 5,065,851 – 5,065,977 |
| Length | 126 |
| Max. P | 0.999726 |

| Location | 5,065,851 – 5,065,944 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 71.17 |
| Mean single sequence MFE | -34.92 |
| Consensus MFE | -9.04 |
| Energy contribution | -9.84 |
| Covariance contribution | 0.80 |
| Combinations/Pair | 1.26 |
| Mean z-score | -3.11 |
| Structure conservation index | 0.26 |
| SVM decision value | 3.59 |
| SVM RNA-class probability | 0.999422 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
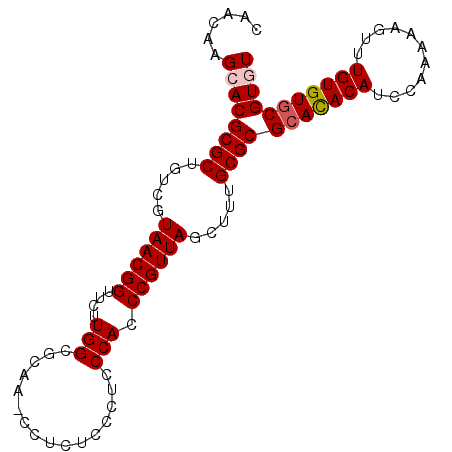
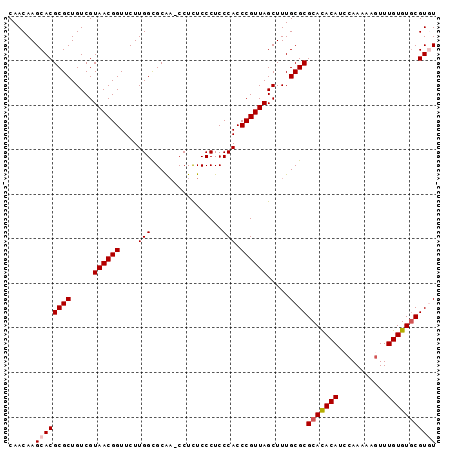
>2R_DroMel_CAF1 5065851 93 + 20766785 ACACAACCACUCCAACC--------AUCCACGAAUAAUC--------AAACGCACACAAACUUUUUGUAUGU---GUGCGCGCAAAGCUAACGGGUGGGAGGGAGA-GG-UUGC------ ...(((((.((((..((--------((((..........--------....(((((((.((.....)).)))---))))((.....))....))))))...)))).-))-))).------ ( -40.20) >DroSec_CAF1 8677 92 + 1 -CACAACCACUCCAACC--------AUCCACGAAUAAUC--------ACACGUACACAAACUUUUUGGAUGU---GUGCGCGCAAAGCUAACGGGUGGGAGGGAGA-GG-UUGC------ -..(((((.((((..((--------((((..........--------....(((((((..(.....)..)))---))))((.....))....))))))...)))).-))-))).------ ( -37.60) >DroSim_CAF1 8822 92 + 1 -CACAACCACUCCAACC--------AUCCACGAAUAAUC--------ACACGCACACAAACUUUUUGGAUGU---GUGCGCGCAAAGCUAACGGGUGGGAGGGAGA-GG-UUGC------ -..(((((.((((..((--------((((..........--------....(((((((..(.....)..)))---))))((.....))....))))))...)))).-))-))).------ ( -39.60) >DroEre_CAF1 8892 93 + 1 ACACAAGCACACCAACC--------AUCCUCGAAUAAUC--------AUACGCACACAAAGUU-UUGGAUGU---AUGCGCGCAAAGCUAACGGGUGGGAGGGAAA-GGGGCGC------ ......((.(.((..((--------((((.........(--------(((((..((.......-.))..)))---))).((.....))....))))))..))....-.).))..------ ( -22.10) >DroAna_CAF1 8258 116 + 1 ACACAUACACUUUGACCCAAGGCUCCCCCUCGAUGGGUUUUUGUUGUUCUCGCACGCAAAGUUAGUGGGGGUAGUGUGG---GAGUGCGAGUGGGAGGGAGGGAGGUAG-UUGGGGUGGU .......(((((..((((....(((((.((((.....((..(..((..(((((((.....))..)))))..))..)..)---)....)))).)))))....)).....)-)..))))).. ( -35.10) >consensus ACACAACCACUCCAACC________AUCCACGAAUAAUC________ACACGCACACAAACUUUUUGGAUGU___GUGCGCGCAAAGCUAACGGGUGGGAGGGAGA_GG_UUGC______ .........((((..((........((((......................(((((((...........)))...))))((.....))....))))))...))))............... ( -9.04 = -9.84 + 0.80)

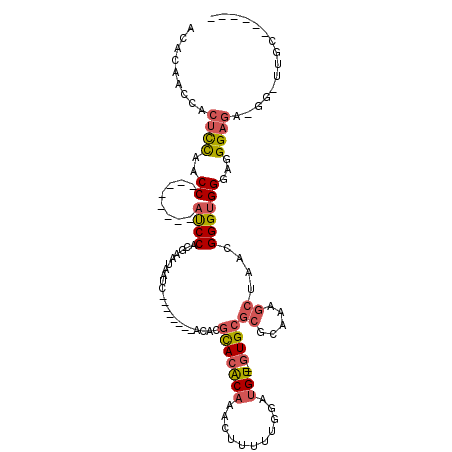
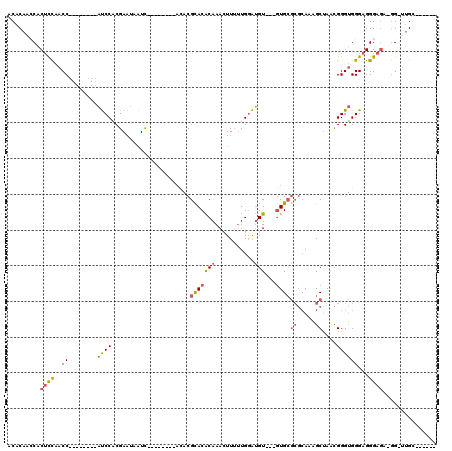
| Location | 5,065,851 – 5,065,944 |
|---|---|
| Length | 93 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 71.17 |
| Mean single sequence MFE | -35.17 |
| Consensus MFE | -8.81 |
| Energy contribution | -11.53 |
| Covariance contribution | 2.72 |
| Combinations/Pair | 1.10 |
| Mean z-score | -3.49 |
| Structure conservation index | 0.25 |
| SVM decision value | 3.95 |
| SVM RNA-class probability | 0.999726 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5065851 93 - 20766785 ------GCAA-CC-UCUCCCUCCCACCCGUUAGCUUUGCGCGCAC---ACAUACAAAAAGUUUGUGUGCGUUU--------GAUUAUUCGUGGAU--------GGUUGGAGUGGUUGUGU ------((((-((-.((((...(((((((......(((.((((((---(((.((.....)).))))))))).)--------)).....)).)).)--------))..)))).)))))).. ( -38.50) >DroSec_CAF1 8677 92 - 1 ------GCAA-CC-UCUCCCUCCCACCCGUUAGCUUUGCGCGCAC---ACAUCCAAAAAGUUUGUGUACGUGU--------GAUUAUUCGUGGAU--------GGUUGGAGUGGUUGUG- ------((((-((-.((((...(((((((......(..((((.((---(((..(.....)..))))).)))).--------.).....)).)).)--------))..)))).)))))).- ( -37.40) >DroSim_CAF1 8822 92 - 1 ------GCAA-CC-UCUCCCUCCCACCCGUUAGCUUUGCGCGCAC---ACAUCCAAAAAGUUUGUGUGCGUGU--------GAUUAUUCGUGGAU--------GGUUGGAGUGGUUGUG- ------((((-((-.((((...(((((((......(..(((((((---(((..(.....)..)))))))))).--------.).....)).)).)--------))..)))).)))))).- ( -44.00) >DroEre_CAF1 8892 93 - 1 ------GCGCCCC-UUUCCCUCCCACCCGUUAGCUUUGCGCGCAU---ACAUCCAA-AACUUUGUGUGCGUAU--------GAUUAUUCGAGGAU--------GGUUGGUGUGCUUGUGU ------((((.((-...((.((((........((...))((((((---(((.....-.....)))))))))..--------........).))).--------))..)).))))...... ( -27.00) >DroAna_CAF1 8258 116 - 1 ACCACCCCAA-CUACCUCCCUCCCUCCCACUCGCACUC---CCACACUACCCCCACUAACUUUGCGUGCGAGAACAACAAAAACCCAUCGAGGGGGAGCCUUGGGUCAAAGUGUAUGUGU ..(((....(-((....(((...(((((.(((((((..---........................)))))))............((.....)))))))....)))....)))....))). ( -28.97) >consensus ______GCAA_CC_UCUCCCUCCCACCCGUUAGCUUUGCGCGCAC___ACAUCCAAAAAGUUUGUGUGCGUGU________GAUUAUUCGUGGAU________GGUUGGAGUGGUUGUGU .................((((((.(((.........(((((((((..................))))))))).........((....))..............))).)))).))...... ( -8.81 = -11.53 + 2.72)

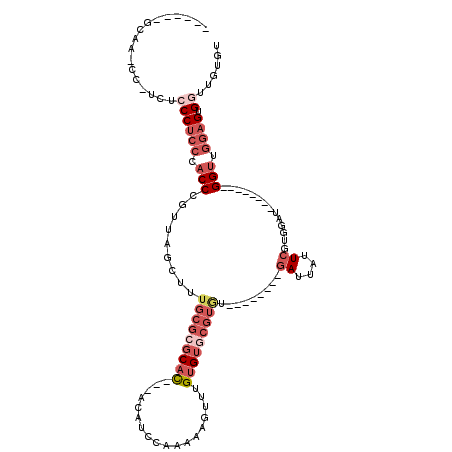
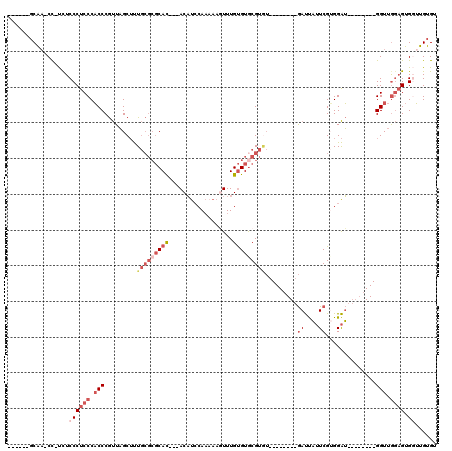
| Location | 5,065,882 – 5,065,977 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 96 |
| Reading direction | reverse |
| Mean pairwise identity | 93.89 |
| Mean single sequence MFE | -26.35 |
| Consensus MFE | -23.85 |
| Energy contribution | -24.42 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.04 |
| Mean z-score | -1.40 |
| Structure conservation index | 0.91 |
| SVM decision value | 0.05 |
| SVM RNA-class probability | 0.557026 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 5065882 95 - 20766785 CAACAAGCACGCGCUGUCGUAACGGUUCUUGGCGCAA-CCUCUCCCUCCCACCCGUUAGCUUUGCGCGCACACAUACAAAAAGUUUGUGUGCGUUU ....((((..((((.....((((((.....((.(...-.....)))......)))))).....))))(((((((.((.....)).))))))))))) ( -28.00) >DroSec_CAF1 8707 95 - 1 CAACAAGCACGCGCUGUCGUAACGGUUCUUGGCGCAA-CCUCUCCCUCCCACCCGUUAGCUUUGCGCGCACACAUCCAAAAAGUUUGUGUACGUGU ......((((((((.....((((((.....((.(...-.....)))......)))))).....))))(.(((((..(.....)..))))).))))) ( -23.70) >DroSim_CAF1 8852 95 - 1 CAACAAGCACGCGCUGUCGUAACGGUUCUUGGCGCAA-CCUCUCCCUCCCACCCGUUAGCUUUGCGCGCACACAUCCAAAAAGUUUGUGUGCGUGU ......((((((((.....((((((.....((.(...-.....)))......)))))).....))))(((((((..(.....)..))))))))))) ( -30.30) >DroEre_CAF1 8923 95 - 1 CAACAAGCACGCGCUGUCGUAACGGUUCUUGGUGCGCCCCUUUCCCUCCCACCCGUUAGCUUUGCGCGCAUACAUCCAA-AACUUUGUGUGCGUAU ......((..((((.....((((((....(((................))).)))))).....))))(((((((.....-.....))))))))).. ( -23.39) >consensus CAACAAGCACGCGCUGUCGUAACGGUUCUUGGCGCAA_CCUCUCCCUCCCACCCGUUAGCUUUGCGCGCACACAUCCAAAAAGUUUGUGUGCGUGU ......((((((((.....((((((....(((................))).)))))).....))))(((((((...........))))))))))) (-23.85 = -24.42 + 0.56)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:01:49 2006