
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,931,930 – 4,932,025 |
| Length | 95 |
| Max. P | 0.895040 |

| Location | 4,931,930 – 4,932,025 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 74.30 |
| Mean single sequence MFE | -28.95 |
| Consensus MFE | -14.93 |
| Energy contribution | -15.18 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.20 |
| Mean z-score | -2.22 |
| Structure conservation index | 0.52 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.895040 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
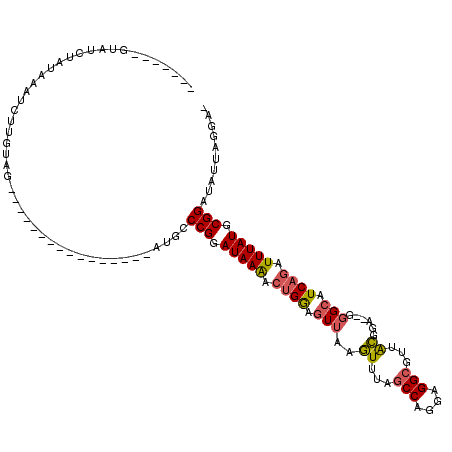
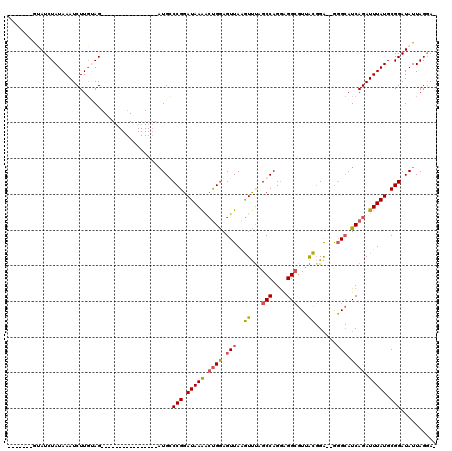
>2R_DroMel_CAF1 4931930 95 + 20766785 -------GUAUCUAUAAAUCUUGUAG----------------AUGCCCGGAUAAAACUGAAGUUAAGUUUAGCCAGAAGGCGUUACGGA--GGGCAUCAGAUUUAUGCGGAUAUUAGGAA -------(((((((((.....)))))----------------))))(((.(((((.((((.(((..((...(((....)))...))...--.))).)))).))))).))).......... ( -30.40) >DroSim_CAF1 162 96 + 1 -------GUAUCUAUAAAUCUUGUAG----------------AUGGCCGGAUAAAACUGGAGUUAAGUUUAGCCAGGAGGCGUUACGGAGAGGGCAUCAGAUUUAUGCGGAUAUUAGGA- -------.((((((((.....)))))----------------))).(((.(((((.(((..(((..((...(((....)))...))......)))..))).))))).))).........- ( -25.40) >DroYak_CAF1 128 94 + 1 -------GAAUCUAUAAAUCUUGGAG----------------AUGCCCGGAUAAAACUGGAGUUAGGUUCAGCCAGGAGGCGUUGCGGA--AGGCAUCAGAUUUAUGCGGAUAUUAGGA- -------..((((((((((((....(----------------(((((.((((..(((....)))..)))).(((....)))........--.))))))))))))))..)))).......- ( -24.90) >DroAna_CAF1 2509 117 + 1 CUCCAACAGAUCUAUAAAUCUUCAAGCGCGCUUUCAAAGAUCCUGGCCGGAUAAGGACGGAGUUAAAUCUAGCCAGGGGGAGUGGUUGG--GCGAAUCAGAUUUAUGCGGAUAUUAGAA- .........((((((((((((.....(((.(..(((....(((((((..(((...(((...)))..)))..)))))))....)))..).--)))....))))))))..)))).......- ( -35.10) >consensus _______GUAUCUAUAAAUCUUGUAG________________AUGCCCGGAUAAAACUGGAGUUAAGUUUAGCCAGGAGGCGUUACGGA__GGGCAUCAGAUUUAUGCGGAUAUUAGGA_ ..............................................(((.(((((.((((.(((..((...(((....)))...))......))).)))).))))).))).......... (-14.93 = -15.18 + 0.25)

| Location | 4,931,930 – 4,932,025 |
|---|---|
| Length | 95 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 74.30 |
| Mean single sequence MFE | -22.52 |
| Consensus MFE | -8.59 |
| Energy contribution | -9.78 |
| Covariance contribution | 1.19 |
| Combinations/Pair | 1.12 |
| Mean z-score | -2.68 |
| Structure conservation index | 0.38 |
| SVM decision value | 0.24 |
| SVM RNA-class probability | 0.650487 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4931930 95 - 20766785 UUCCUAAUAUCCGCAUAAAUCUGAUGCCC--UCCGUAACGCCUUCUGGCUAAACUUAACUUCAGUUUUAUCCGGGCAU----------------CUACAAGAUUUAUAGAUAC------- .......((((...(((((((((((((((--........(((....))).(((((.......))))).....))))))----------------)....)))))))).)))).------- ( -24.60) >DroSim_CAF1 162 96 - 1 -UCCUAAUAUCCGCAUAAAUCUGAUGCCCUCUCCGUAACGCCUCCUGGCUAAACUUAACUCCAGUUUUAUCCGGCCAU----------------CUACAAGAUUUAUAGAUAC------- -......((((...((((((((((((.((..........(((....))).(((((.......))))).....)).)))----------------)....)))))))).)))).------- ( -17.50) >DroYak_CAF1 128 94 - 1 -UCCUAAUAUCCGCAUAAAUCUGAUGCCU--UCCGCAACGCCUCCUGGCUGAACCUAACUCCAGUUUUAUCCGGGCAU----------------CUCCAAGAUUUAUAGAUUC------- -.............(((((((((((((((--........(((....)))((((.((......)).))))...))))))----------------)....))))))))......------- ( -19.80) >DroAna_CAF1 2509 117 - 1 -UUCUAAUAUCCGCAUAAAUCUGAUUCGC--CCAACCACUCCCCCUGGCUAGAUUUAACUCCGUCCUUAUCCGGCCAGGAUCUUUGAAAGCGCGCUUGAAGAUUUAUAGAUCUGUUGGAG -.(((((((.....((((((((....(((--............((((((..(((..............)))..))))))............))).....)))))))).....))))))). ( -28.19) >consensus _UCCUAAUAUCCGCAUAAAUCUGAUGCCC__UCCGCAACGCCUCCUGGCUAAACUUAACUCCAGUUUUAUCCGGCCAU________________CUACAAGAUUUAUAGAUAC_______ ..........(((.(((((.(((.(((.......)))..(((....)))............))).))))).))).............................................. ( -8.59 = -9.78 + 1.19)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 10:00:28 2006