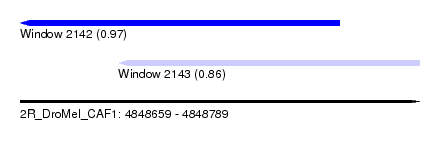

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,848,659 – 4,848,789 |
| Length | 130 |
| Max. P | 0.965181 |

| Location | 4,848,659 – 4,848,763 |
|---|---|
| Length | 104 |
| Sequences | 4 |
| Columns | 109 |
| Reading direction | reverse |
| Mean pairwise identity | 79.34 |
| Mean single sequence MFE | -23.98 |
| Consensus MFE | -15.21 |
| Energy contribution | -14.78 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.36 |
| Structure conservation index | 0.63 |
| SVM decision value | 1.58 |
| SVM RNA-class probability | 0.965181 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
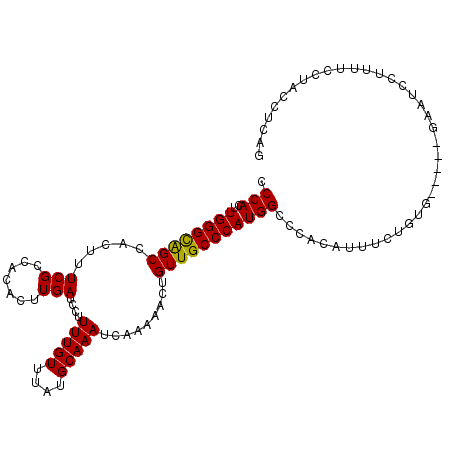
>2R_DroMel_CAF1 4848659 104 - 20766785 CCCACUUGGGUAGCCACUUUCGCCACACUUGAUCCUUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCCCACAUUUCUGUG-----GUAUCCUUUUCCUACCUCAG ....((.(((((((((.....((.(((.((......(((((....))))).....)).))).))...))))(((((....))))-----)...........))))).)) ( -21.40) >DroGri_CAF1 20024 81 - 1 CCCACUUGGGUGGCCACUUUCGGUACACUUGAACC-UUUGUUUAUGCAAAUCAA-AACGGUUGCCCAUGGCCCACUUUG----------UC---GC------------- .(((..((((..(((......(((.(....).)))-(((((....)))))....-...)))..))))))).........----------..---..------------- ( -22.00) >DroEre_CAF1 5280 109 - 1 CCCACUUGGGCAGCUACUUUCGCCACACUUGAUCCCUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCCCACAUUUCAGUGGGUAGGAAUCCUUUUCCUACCUCAG .(((..((((((((.....(((.......)))....(((((....))))).........)))))))))))..(((......)))((((((((.....)))))))).... ( -30.60) >DroYak_CAF1 9516 104 - 1 CCCACUUGGGCAGCCACUUUCGCCACACUUGAUCCUUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCCCACAUUUCUGUG-----GAAUCAUUUUCCUACCUCAG .(((..((((((((.....(((.......)))....(((((....))))).........))))))))))).(((((....))))-----)................... ( -21.90) >consensus CCCACUUGGGCAGCCACUUUCGCCACACUUGAUCCUUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCCCACAUUUCUGUG_____GAAUCCUUUUCCUACCUCAG .(((..((((((((.....(((.......)))....(((((....))))).........)))))))))))....................................... (-15.21 = -14.78 + -0.44)


| Location | 4,848,691 – 4,848,789 |
|---|---|
| Length | 98 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | reverse |
| Mean pairwise identity | 79.87 |
| Mean single sequence MFE | -24.35 |
| Consensus MFE | -11.04 |
| Energy contribution | -11.82 |
| Covariance contribution | 0.78 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.34 |
| Structure conservation index | 0.45 |
| SVM decision value | 0.81 |
| SVM RNA-class probability | 0.856702 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
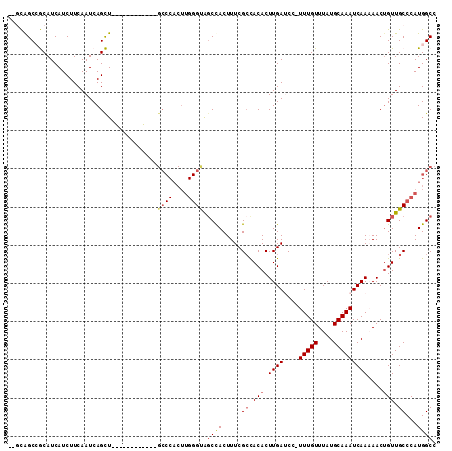
>2R_DroMel_CAF1 4848691 98 - 20766785 --GCAGCCGCAUCAUCUUCAAUCAGCU------------GCCCACUUGGGUAGCCACUUUCGCCACACUUGAUCCUUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCC --...((((...............(((------------((((....))))))).......((.(((.((......(((((....))))).....)).))).))...)))). ( -23.40) >DroGri_CAF1 20035 109 - 1 UUGUGGCAACAUCAUCUUCAAUCAGUU-AGCCCUAUCAUGCCCACUUGGGUGGCCACUUUCGGUACACUUGAACC-UUUGUUUAUGCAAAUCAA-AACGGUUGCCCAUGGCC ....((((((..............)))-.))).......(((....((((..(((......(((.(....).)))-(((((....)))))....-...)))..)))).))). ( -27.44) >DroEre_CAF1 5317 98 - 1 --ACAGCCGCAUCAUCUUCAAUCAGCU------------GCCCACUUGGGCAGCUACUUUCGCCACACUUGAUCCCUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCC --...((((..............((((------------((((....))))))))......((.(((.(((((....((((....)))))))))....))).))...)))). ( -26.80) >DroYak_CAF1 9548 98 - 1 --ACAGCCGCAUCAUCUUCAAUCAGCU------------GCCCACUUGGGCAGCCACUUUCGCCACACUUGAUCCUUUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCC --...((((...............(((------------((((....))))))).......((.(((.((......(((((....))))).....)).))).))...)))). ( -25.40) >DroMoj_CAF1 27328 100 - 1 CUGUGGCAACAUCAUCUUCAAUCAGUUUGGCCCUAUACU----------GUGCCCACUUUCGCUACACUUGAACC-UUUGUUUAUGCAAAUCAA-AACUGUUGCCCAUGGCC (((((((((((............(((..(((.(......----------).))).)))..........((((...-(((((....)))))))))-...))))).)))))).. ( -18.70) >consensus __GCAGCCGCAUCAUCUUCAAUCAGCU____________GCCCACUUGGGUAGCCACUUUCGCCACACUUGAUCC_UUUGUUUAUGCAAAUCAAAAACUGUUGCCCAUGGCC .......................................((((....)))).((((.....((.(((.((((....(((((....)))))))))....))).))...)))). (-11.04 = -11.82 + 0.78)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:59:39 2006