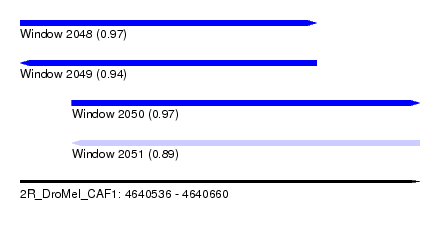
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,640,536 – 4,640,660 |
| Length | 124 |
| Max. P | 0.973345 |

| Location | 4,640,536 – 4,640,628 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | forward |
| Mean pairwise identity | 88.41 |
| Mean single sequence MFE | -31.43 |
| Consensus MFE | -27.65 |
| Energy contribution | -27.67 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.20 |
| Mean z-score | -3.04 |
| Structure conservation index | 0.88 |
| SVM decision value | 1.71 |
| SVM RNA-class probability | 0.973345 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4640536 92 + 20766785 UUUUAUUUGCCCCGGCUACUCAUUGCCUGGCCUUGCUCAUCGCGCGUUGCCCAAUGUUACGCAUGAUGAGUUGUUGAUACUUGGCUGUACAG ........((((((((........))).))....((((((((.((((.((.....)).)))).))))))))...........)))....... ( -26.50) >DroEre_CAF1 12113 92 + 1 UUGUAUUUGCUUUGACUACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAACGUUAUGCAUGAUGAGUUGUUAAUACUUGGCAGUGCAG ..................(((((((((((((...(((((((((((((.((.....)).))))))))))))).))))......))))))).)) ( -30.70) >DroYak_CAF1 11732 92 + 1 UUGUAUUUGCUCCGGCUACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAAUGCUUCGCAUGAUGAGUGGUUAAUACCUGGCUGUGCAG .(((((.......(((........))).((((..((((((((((((..((.....))..))))))))))))(((....))).))))))))). ( -37.10) >consensus UUGUAUUUGCUCCGGCUACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAAUGUUACGCAUGAUGAGUUGUUAAUACUUGGCUGUGCAG .(((((..(((..(((........)))((((...(((((((((((((.((.....)).))))))))))))).))))......))).))))). (-27.65 = -27.67 + 0.01)

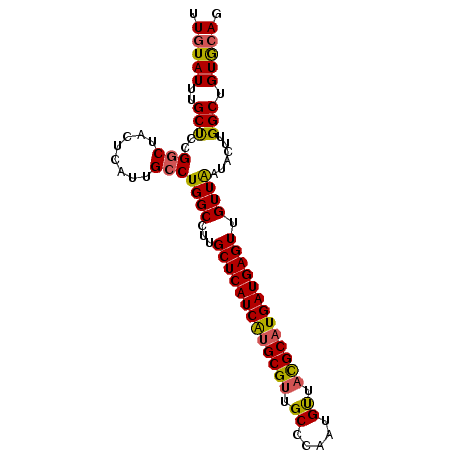
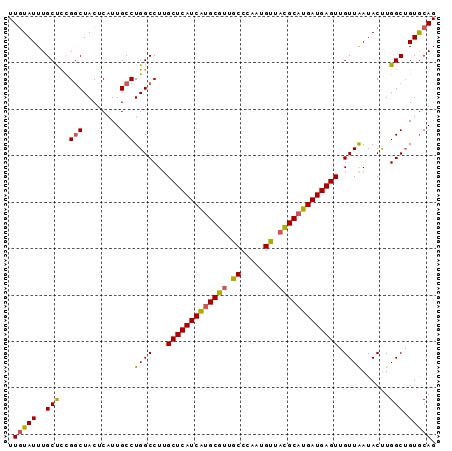
| Location | 4,640,536 – 4,640,628 |
|---|---|
| Length | 92 |
| Sequences | 3 |
| Columns | 92 |
| Reading direction | reverse |
| Mean pairwise identity | 88.41 |
| Mean single sequence MFE | -27.97 |
| Consensus MFE | -23.85 |
| Energy contribution | -23.85 |
| Covariance contribution | 0.01 |
| Combinations/Pair | 1.14 |
| Mean z-score | -2.77 |
| Structure conservation index | 0.85 |
| SVM decision value | 1.29 |
| SVM RNA-class probability | 0.938765 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4640536 92 - 20766785 CUGUACAGCCAAGUAUCAACAACUCAUCAUGCGUAACAUUGGGCAACGCGCGAUGAGCAAGGCCAGGCAAUGAGUAGCCGGGGCAAAUAAAA .......(((............((((((.(((((...........))))).)))))).....((.(((........))))))))........ ( -24.90) >DroEre_CAF1 12113 92 - 1 CUGCACUGCCAAGUAUUAACAACUCAUCAUGCAUAACGUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUAGUCAAAGCAAAUACAA .(((.(((((..((....))..((((((((((.....(((....)))))))))))))...)).))))))....................... ( -23.20) >DroYak_CAF1 11732 92 - 1 CUGCACAGCCAGGUAUUAACCACUCAUCAUGCGAAGCAUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUAGCCGGAGCAAAUACAA .(((...(((.(((....))).(((((((((((..((.....))..)))))))))))...)))..(((........)))...)))....... ( -35.80) >consensus CUGCACAGCCAAGUAUUAACAACUCAUCAUGCGUAACAUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUAGCCGGAGCAAAUACAA .(((...(((..((....))..(((((((((((.............)))))))))))...)))..(((........)))...)))....... (-23.85 = -23.85 + 0.01)

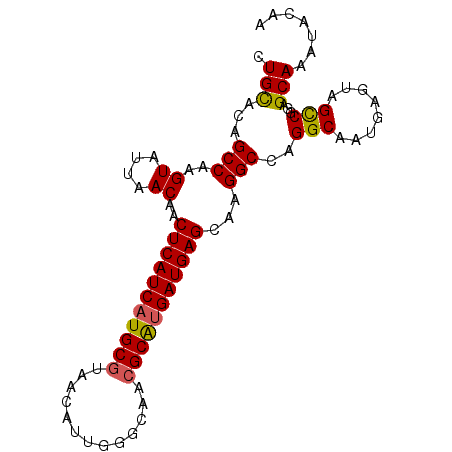
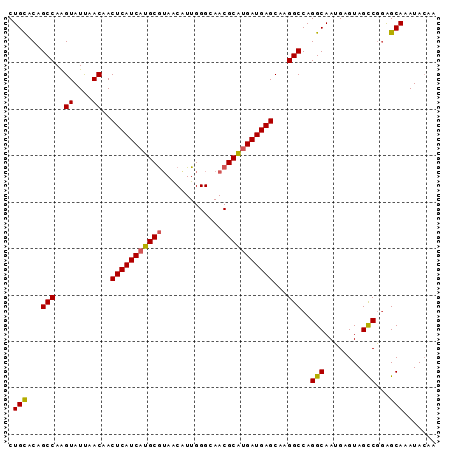
| Location | 4,640,552 – 4,640,660 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 115 |
| Reading direction | forward |
| Mean pairwise identity | 85.42 |
| Mean single sequence MFE | -35.07 |
| Consensus MFE | -31.88 |
| Energy contribution | -32.33 |
| Covariance contribution | 0.45 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.32 |
| Structure conservation index | 0.91 |
| SVM decision value | 1.62 |
| SVM RNA-class probability | 0.968096 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
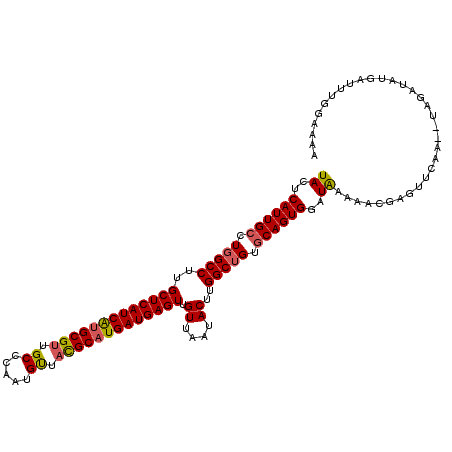
>2R_DroMel_CAF1 4640552 108 + 20766785 UACUCAUUGCCUGGCCUUGCUCAUCGCGCGUUGCCCAAUGUUACGCAUGAUGAGUUGUUGAUACUUGGCUGUACAGUGGAUAAAAACGAGUUCAA--UUGAUGUGAUAUG----- ..(.(((((..((..(((((((((((.((((.((.....)).)))).)))))))(..(((.(((......))))))..).......))))..)).--.))))).).....----- ( -27.90) >DroEre_CAF1 12129 115 + 1 UACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAACGUUAUGCAUGAUGAGUUGUUAAUACUUGGCAGUGCAGUGGAUGAAAACGAGUUCAGUAUAUAUAUGAUUUGGAAAA (((((((((((((((...(((((((((((((.((.....)).))))))))))))).))))......))))))).))))........(((((.((.........)))))))..... ( -35.60) >DroYak_CAF1 11748 113 + 1 UACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAAUGCUUCGCAUGAUGAGUGGUUAAUACCUGGCUGUGCAGUGGAUAGAAACGAGUUCAA--UAGAUAUGAUUUGGAAAA ((..((((((.(((((..((((((((((((..((.....))..))))))))))))(((....))).))))).))))))..))....(((((.((.--......)))))))..... ( -41.70) >consensus UACUCAUUGCCUGGCCUUGCUCAUCAUGCGUUGCCCAAUGUUACGCAUGAUGAGUUGUUAAUACUUGGCUGUGCAGUGGAUAAAAACGAGUUCAA__UAGAUAUGAUUUGGAAAA ((..((((((.(((((..(((((((((((((.((.....)).))))))))))))).((....))..))))).))))))..))................................. (-31.88 = -32.33 + 0.45)

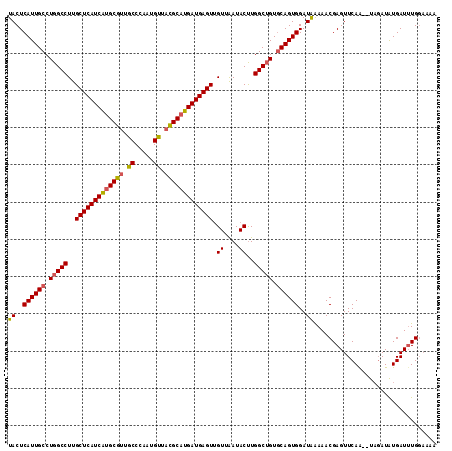
| Location | 4,640,552 – 4,640,660 |
|---|---|
| Length | 108 |
| Sequences | 3 |
| Columns | 115 |
| Reading direction | reverse |
| Mean pairwise identity | 85.42 |
| Mean single sequence MFE | -28.77 |
| Consensus MFE | -24.79 |
| Energy contribution | -25.02 |
| Covariance contribution | 0.23 |
| Combinations/Pair | 1.08 |
| Mean z-score | -2.15 |
| Structure conservation index | 0.86 |
| SVM decision value | 0.98 |
| SVM RNA-class probability | 0.894021 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4640552 108 - 20766785 -----CAUAUCACAUCAA--UUGAACUCGUUUUUAUCCACUGUACAGCCAAGUAUCAACAACUCAUCAUGCGUAACAUUGGGCAACGCGCGAUGAGCAAGGCCAGGCAAUGAGUA -----.............--....((((((..........(((...(((..((....))..((((((.(((((...........))))).))))))...)))...))))))))). ( -23.30) >DroEre_CAF1 12129 115 - 1 UUUUCCAAAUCAUAUAUAUACUGAACUCGUUUUCAUCCACUGCACUGCCAAGUAUUAACAACUCAUCAUGCAUAACGUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUA ........................((((((..........(((.(((((..((....))..((((((((((.....(((....)))))))))))))...)).)))))))))))). ( -26.90) >DroYak_CAF1 11748 113 - 1 UUUUCCAAAUCAUAUCUA--UUGAACUCGUUUCUAUCCACUGCACAGCCAGGUAUUAACCACUCAUCAUGCGAAGCAUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUA .........(((......--.)))((((((..........(((...(((.(((....))).(((((((((((..((.....))..)))))))))))...)))...))))))))). ( -36.10) >consensus UUUUCCAAAUCAUAUCUA__UUGAACUCGUUUUUAUCCACUGCACAGCCAAGUAUUAACAACUCAUCAUGCGUAACAUUGGGCAACGCAUGAUGAGCAAGGCCAGGCAAUGAGUA ........................((((((..........(((...(((..((....))..(((((((((((.............)))))))))))...)))...))))))))). (-24.79 = -25.02 + 0.23)

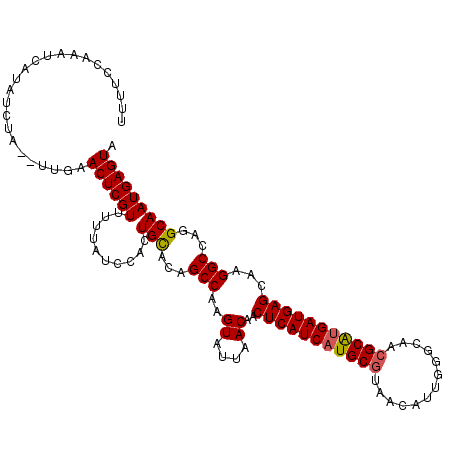
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:58:11 2006