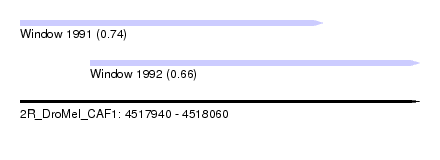

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,517,940 – 4,518,060 |
| Length | 120 |
| Max. P | 0.739160 |

| Location | 4,517,940 – 4,518,031 |
|---|---|
| Length | 91 |
| Sequences | 5 |
| Columns | 91 |
| Reading direction | forward |
| Mean pairwise identity | 91.43 |
| Mean single sequence MFE | -25.04 |
| Consensus MFE | -20.60 |
| Energy contribution | -21.12 |
| Covariance contribution | 0.52 |
| Combinations/Pair | 1.11 |
| Mean z-score | -1.49 |
| Structure conservation index | 0.82 |
| SVM decision value | 0.44 |
| SVM RNA-class probability | 0.739160 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
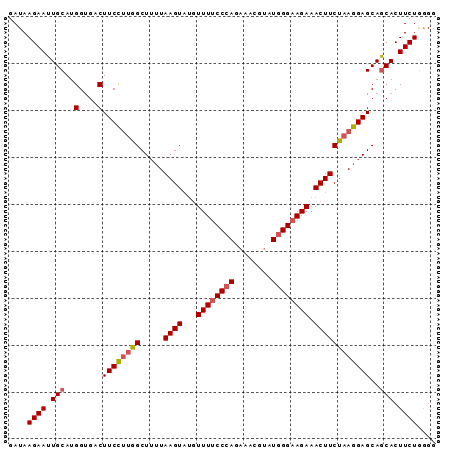
>2R_DroMel_CAF1 4517940 91 + 20766785 GAUAAGAAUUGCAUGGUGACUUCCUUGGCUUUUAAGUACGUUUUCCGAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGGGG ....((((.(((..(....)((((((((((((...(((((((((....)))))))))..))))......))))))))..))).)))).... ( -25.00) >DroSec_CAF1 52305 91 + 1 GAUAAGAAUUGCAUGGUGACUUCCUUGGCUUUUAAGUAUCUUUUCCCAGAAACGAAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGAGG ....((((.(((..(....)((((((((.....((((.(((..(((((........)))))))).))))))))))))..))).)))).... ( -26.80) >DroSim_CAF1 51232 91 + 1 GAUAAGAAUUGCAUGGUGACUUCCUUGGCUUUUAAGUAUGUUUUCCCAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUAGGG ....((((.(((..(....)((((((((.....((((...((((((((........)))))))).))))))))))))..))).)))).... ( -27.60) >DroEre_CAF1 51422 91 + 1 GAUAAGAAUUGCAUGGCGACUUCUUCGGCUUUUAAGUGUGUUUUCCCAGAAAUGUGUGGGAAGAAACUUCCAAGGAGCAACACUUCUGGGG ...(((((((((...)))).)))))...................(((((((.(((((.(((((...))))).....)).))).))))))). ( -23.20) >DroYak_CAF1 53888 89 + 1 GAUAAGAAUUGCAUGGUGACUUCCGUGGCU--UAAGUAUGUUUUCCCAGAAAUUUAUGGGUAGAAACUUCUAAGGAGCAGCACUUCUGGGG ....((((.(((((((......)))).(((--(.((...(((((((((........))))..)))))..))...)))).))).)))).... ( -22.60) >consensus GAUAAGAAUUGCAUGGUGACUUCCUUGGCUUUUAAGUAUGUUUUCCCAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGGGG ....((((.(((...((....(((((((.....((((...((((((((........)))))))).))))))))))))).))).)))).... (-20.60 = -21.12 + 0.52)


| Location | 4,517,961 – 4,518,060 |
|---|---|
| Length | 99 |
| Sequences | 5 |
| Columns | 110 |
| Reading direction | forward |
| Mean pairwise identity | 84.72 |
| Mean single sequence MFE | -27.76 |
| Consensus MFE | -19.82 |
| Energy contribution | -19.58 |
| Covariance contribution | -0.24 |
| Combinations/Pair | 1.17 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.71 |
| SVM decision value | 0.27 |
| SVM RNA-class probability | 0.659913 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
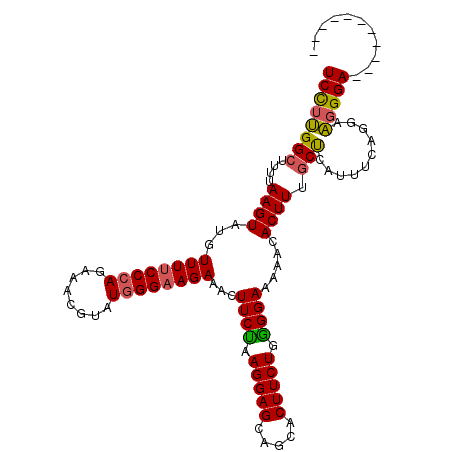
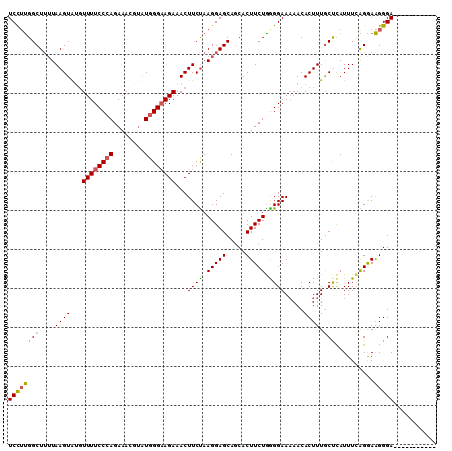
>2R_DroMel_CAF1 4517961 99 + 20766785 UCCUUGGCUUUUAAGUACGUUUUCCGAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGGGGAAAAACACUUUGCUCAUUUCAGGAAGGGA----------- (((((.((((((..(((((((((....))))))))).(((((...))))).))))))........((((.(((.....))).)))).......))))).----------- ( -23.30) >DroSec_CAF1 52326 99 + 1 UCCUUGGCUUUUAAGUAUCUUUUCCCAGAAACGAAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGAGGAAAAACACUUUACCCAUUUCAGAAAGGGA----------- (((((.((((((.((....((((((((........)))))))).....)).)))))).....((((((((.(((....))).))....)))))))))))----------- ( -25.40) >DroSim_CAF1 51253 99 + 1 UCCUUGGCUUUUAAGUAUGUUUUCCCAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUAGGGAAAAACACUUUGCCCAUUUCAGAAAGGGA----------- (((((.((((((.((....((((((((........)))))))).....)).))))))..........(((.(((....))).)))........))))).----------- ( -25.70) >DroEre_CAF1 51443 99 + 1 UCUUCGGCUUUUAAGUGUGUUUUCCCAGAAAUGUGUGGGAAGAAACUUCCAAGGAGCAACACUUCUGGGGAAAAACACUUUGCUUAUUUUGGGAGGGGA----------- ((((((((....((((((.(((((((((((.(((((.(((((...))))).....)).))).))))))))))).)))))).)))........)))))..----------- ( -35.00) >DroYak_CAF1 53909 108 + 1 UCCGUGGCU--UAAGUAUGUUUUCCCAGAAAUUUAUGGGUAGAAACUUCUAAGGAGCAGCACUUCUGGGGAACAACACUUUGCUCAUUUUGGGAAGGGAAAUUUAGCUUU .....((((--.((((.(.((((((((((.....(((((((((...((((.(((((.....)))))..))))......))))))))))))))))))).).)))))))).. ( -29.40) >consensus UCCUUGGCUUUUAAGUAUGUUUUCCCAGAAACGUAUGGGAAGAAACUUCUAAGGAGCAGCACUUCUGGGGAAAAACACUUUGCUCAUUUCAGGAAGGGA___________ ((((((((....((((...((((((((........))))))))...((((.(((((.....))))).)))).....)))).)))..........)))))........... (-19.82 = -19.58 + -0.24)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:57:14 2006