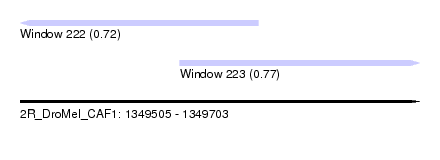
| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 1,349,505 – 1,349,703 |
| Length | 198 |
| Max. P | 0.773536 |

| Location | 1,349,505 – 1,349,623 |
|---|---|
| Length | 118 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 88.33 |
| Mean single sequence MFE | -21.01 |
| Consensus MFE | -16.36 |
| Energy contribution | -16.37 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.05 |
| Mean z-score | -1.69 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.39 |
| SVM RNA-class probability | 0.716593 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
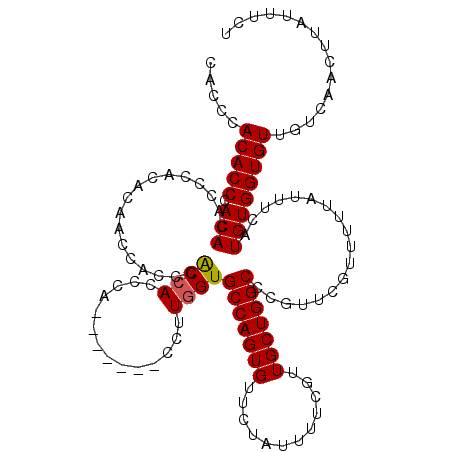
>2R_DroMel_CAF1 1349505 118 - 20766785 CACCCACACCCACAUAGCCACC-CACCACUCGCCACCCAG-CACUCCCUUGGUGCCAGUGUUCUAUUUUCGUUGCUGGCCCGUUCGUUUUUAUUUCAUGUGGUGUUGUCAACUUAUUUCU ......................-((((((..((......)-)........((.(((((((............))))))))).................))))))................ ( -22.40) >DroSec_CAF1 67749 113 - 1 CACCCACACCCACACACCCACACAACCACCCGCCACCCA-------CUUUGGUGCCAGUGUUCUAUUUUCGUUGCUGGCCCGUUCGUUUUUAUUUCAUGUGGUGUUGUCAACUUAUUUCU ...........(((((((.(((.........((((....-------...))))(((((((............)))))))..................))))))).)))............ ( -19.50) >DroSim_CAF1 45884 111 - 1 CACCCACACCCACACACCCA--CAACCACCCACCACCCA-------CUUUGGUGCCAGUGUUCUAUUUUCGUUGCUGGCCCGUUCGUUUUUAUUUCAUGUGGUGUUGUCAACUUAUUUCU ...........(((((((.(--((.......((((....-------...))))(((((((............)))))))..................))))))).)))............ ( -19.50) >DroYak_CAF1 35862 120 - 1 CACACACACCCACACACCAACACUGCCACCGACAACCCACUUACUCCCUUAGUGCCAGUGCUCUAUUUUCGUUGCUGGCCCGUUCGUUUUUAUUUCAUGUGGUGUUGUCAACUUAUUUCU ..............................((((((((((..(((.....)))((((((..............))))))...................)))).))))))........... ( -22.64) >consensus CACCCACACCCACACACCCACACAACCACCCACCACCCA_______CCUUGGUGCCAGUGUUCUAUUUUCGUUGCUGGCCCGUUCGUUUUUAUUUCAUGUGGUGUUGUCAACUUAUUUCU .....(((((.(((.................((((..............))))(((((((............)))))))..................))))))))............... (-16.36 = -16.37 + 0.00)

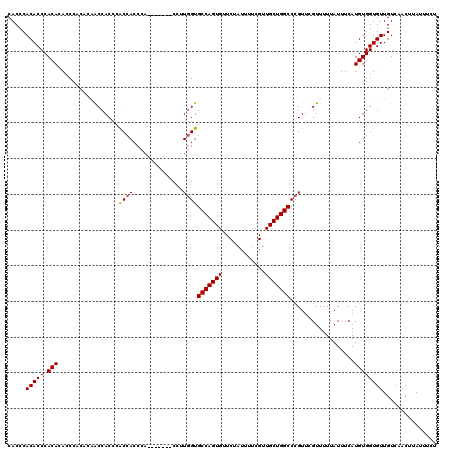
| Location | 1,349,584 – 1,349,703 |
|---|---|
| Length | 119 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 92.35 |
| Mean single sequence MFE | -35.75 |
| Consensus MFE | -28.20 |
| Energy contribution | -28.95 |
| Covariance contribution | 0.75 |
| Combinations/Pair | 1.09 |
| Mean z-score | -2.04 |
| Structure conservation index | 0.79 |
| SVM decision value | 0.54 |
| SVM RNA-class probability | 0.773536 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
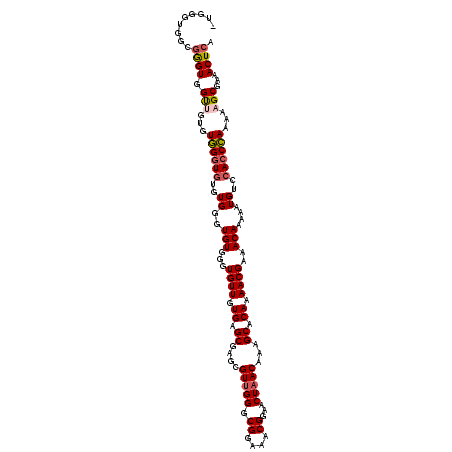
>2R_DroMel_CAF1 1349584 119 + 20766785 CUGGGUGGCGAGUGGUG-GGUGGCUAUGUGGGUGUGGGUGUUGUGAGCGAGCGUUGGGCGGAAACGGAACUAACAAAGCACAAAACGAAACAAAAAUGUCCACCCAAAAAGCGAAACUCA ..((((..((..(..((-(((((.(((.((..(((..(((((((......))(((((.((....))...)))))..)))))...)))...))...))).)))))))..)..))..)))). ( -41.40) >DroSec_CAF1 67823 119 + 1 -UGGGUGGCGGGUGGUUGUGUGGGUGUGUGGGUGUGGGUGUUGUGAGCGAGCGUUGGGCGGAAACGGAACUAACAAAGCACAAAACGAAACAAAAAUGUCCACCCAAAAAGCGAAACUCA -........((((..((((.((((((..((..(((...((((.((.((....(((((.((....))...)))))...)).)).))))..)))....))..))))))....)))).)))). ( -35.50) >DroSim_CAF1 45958 117 + 1 -UGGGUGGUGGGUGGUUG--UGGGUGUGUGGGUGUGGGUGUUGUGAGCGAGCGUUGGGCGGAAACGGAACUAACAAAGCACAAAACGAAACAAAAAUGUCCACCCAAAAAGCGAAACUCA -.((((..(((((((.((--(.....(((...(((..(((((((......))(((((.((....))...)))))..)))))...)))..)))...))).))))))).........)))). ( -33.80) >DroYak_CAF1 35942 120 + 1 GUGGGUUGUCGGUGGCAGUGUUGGUGUGUGGGUGUGUGUGUUGUGAGCGAGCGUCGGGCGGAAACGGAACUAACAAAGCACAAAACGAAACAAAAAUGUCCACCAAAAAAGCGAAACUCA .((((((.(((((((((...(((...(((...(((((...((((.((((.....))..((....))...)).)))).)))))..)))...)))...)).)))))........)))))))) ( -32.30) >consensus _UGGGUGGCGGGUGGUUGUGUGGGUGUGUGGGUGUGGGUGUUGUGAGCGAGCGUUGGGCGGAAACGGAACUAACAAAGCACAAAACGAAACAAAAAUGUCCACCCAAAAAGCGAAACUCA .........((((.(((...((((((..((..(((...((((.((.((....(((((.((....))...)))))...)).)).))))..)))....))..))))))...)))...)))). (-28.20 = -28.95 + 0.75)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:27:21 2006