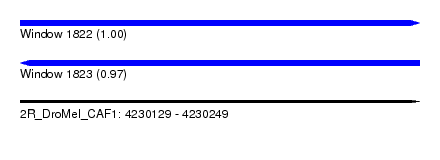

| Sequence ID | 2R_DroMel_CAF1 |
|---|---|
| Location | 4,230,129 – 4,230,249 |
| Length | 120 |
| Max. P | 0.995374 |

| Location | 4,230,129 – 4,230,249 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 95.17 |
| Mean single sequence MFE | -32.30 |
| Consensus MFE | -29.06 |
| Energy contribution | -29.18 |
| Covariance contribution | 0.12 |
| Combinations/Pair | 1.09 |
| Mean z-score | -1.97 |
| Structure conservation index | 0.90 |
| SVM decision value | 2.57 |
| SVM RNA-class probability | 0.995374 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4230129 120 + 20766785 ACAGAUUGUUGCAGCUAAAAUCGGGUCAGAUGCGGCCGAAAAGGAAUUGAUGCUUUCCUCGCCAGCAUCACAAUAUGUUUUUGGCUUUAAUGCUUUUGUUUACUAAAUUAGCAGCAACUA .......(((((.(((((.....(((((((.(((((((((((.(.(((((((((.........)))))))..)).).))))))))).....)).))))...)))...))))).))))).. ( -32.30) >DroSec_CAF1 42041 120 + 1 ACAGAUUGUUGCAGCUAAGACCGGGUCAGAUGCGGCCGAAAAGGAAUUGAUGCUUUCCUUGCCAGCAUCACAAUAUGUUUUUGGUUUUAAUGCUUUUGUUUACUAAAUAAGCAGCAACUA .......((((((((((((((((((...((((((((....((((((........))))))))).)))))((.....)).))))))))))..)))..((((((.....))))))))))).. ( -34.80) >DroSim_CAF1 73231 120 + 1 ACAGAUUGUUGCAGCUAAGACCGGGUCAGAUGCGGCCGAAAAGGAAUUGAUGCUUUCCUUGCCAGCAUCACAAUAUGUUUUUGGCUUUAAUGCUUUUGUUUACUAAAUAAGCAGCAACUA .......(((((((((((((.((.....((((((((....((((((........))))))))).)))))......)).))))))))....(((((...((....))..)))))))))).. ( -34.60) >DroEre_CAF1 43330 120 + 1 ACAGAUUGUUGCAGCUAUGAUCGGGUCAGAUGCGGCCGAAAAGGAAUUGAUGCUUUCCUCGCCAGCAUCACAAUAUGUUUUUGGCUUUAAUGCUUUUCUUUACUAAAUUAGCAGCAACUA .......(((((.((((......(((.(((.(((((((((((.(.(((((((((.........)))))))..)).).))))))))).....))...)))..)))....)))).))))).. ( -31.50) >DroYak_CAF1 45419 120 + 1 ACAGAUUGUUGCAGCUAAGAUCAAGUCAGAUGCGGCCGAAAAAUAAUUGAUUCGUUCCUCGCCAGCAUCACAAUAUGUUUUUGGCUUUAAUGCUUUUGUUUACUAAAUUAGCAGCAACUA .......(((((((((((((.((.((..((((((((.((..((..(.....)..))..))))).)))))...)).)).))))))))....((((...(((.....))).))))))))).. ( -28.30) >consensus ACAGAUUGUUGCAGCUAAGAUCGGGUCAGAUGCGGCCGAAAAGGAAUUGAUGCUUUCCUCGCCAGCAUCACAAUAUGUUUUUGGCUUUAAUGCUUUUGUUUACUAAAUUAGCAGCAACUA .......(((((((((((((.((.....((((((((.....(((((........))))).))).)))))......)).))))))))....((((...............))))))))).. (-29.06 = -29.18 + 0.12)

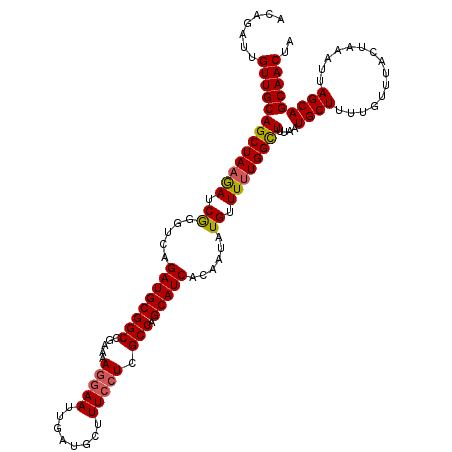
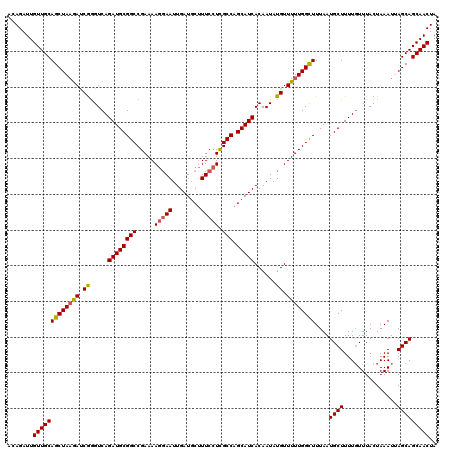
| Location | 4,230,129 – 4,230,249 |
|---|---|
| Length | 120 |
| Sequences | 5 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 95.17 |
| Mean single sequence MFE | -29.58 |
| Consensus MFE | -25.80 |
| Energy contribution | -26.44 |
| Covariance contribution | 0.64 |
| Combinations/Pair | 1.03 |
| Mean z-score | -1.78 |
| Structure conservation index | 0.87 |
| SVM decision value | 1.65 |
| SVM RNA-class probability | 0.969835 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2R_DroMel_CAF1 4230129 120 - 20766785 UAGUUGCUGCUAAUUUAGUAAACAAAAGCAUUAAAGCCAAAAACAUAUUGUGAUGCUGGCGAGGAAAGCAUCAAUUCCUUUUCGGCCGCAUCUGACCCGAUUUUAGCUGCAACAAUCUGU ..(((((.(((((..............((......))..............(((((.(((((((((........))))))....))))))))..........))))).)))))....... ( -30.10) >DroSec_CAF1 42041 120 - 1 UAGUUGCUGCUUAUUUAGUAAACAAAAGCAUUAAAACCAAAAACAUAUUGUGAUGCUGGCAAGGAAAGCAUCAAUUCCUUUUCGGCCGCAUCUGACCCGGUCUUAGCUGCAACAAUCUGU ..(((((((((.....))))......(((..(((.(((....((.....))(((((.(((((((((........))))))....))))))))......))).)))))))))))....... ( -29.00) >DroSim_CAF1 73231 120 - 1 UAGUUGCUGCUUAUUUAGUAAACAAAAGCAUUAAAGCCAAAAACAUAUUGUGAUGCUGGCAAGGAAAGCAUCAAUUCCUUUUCGGCCGCAUCUGACCCGGUCUUAGCUGCAACAAUCUGU ..(((((((((.....))))......(((..(((..((....((.....))(((((.(((((((((........))))))....))))))))......))..)))))))))))....... ( -29.70) >DroEre_CAF1 43330 120 - 1 UAGUUGCUGCUAAUUUAGUAAAGAAAAGCAUUAAAGCCAAAAACAUAUUGUGAUGCUGGCGAGGAAAGCAUCAAUUCCUUUUCGGCCGCAUCUGACCCGAUCAUAGCUGCAACAAUCUGU ..(((((.((((...............((......))..............(((((.(((((((((........))))))....))))))))...........)))).)))))....... ( -29.50) >DroYak_CAF1 45419 120 - 1 UAGUUGCUGCUAAUUUAGUAAACAAAAGCAUUAAAGCCAAAAACAUAUUGUGAUGCUGGCGAGGAACGAAUCAAUUAUUUUUCGGCCGCAUCUGACUUGAUCUUAGCUGCAACAAUCUGU ..(((((.(((((...(((........((......))..............(((((.(.(((((((..(.....)..))))))).).)))))..))).....))))).)))))....... ( -29.60) >consensus UAGUUGCUGCUAAUUUAGUAAACAAAAGCAUUAAAGCCAAAAACAUAUUGUGAUGCUGGCGAGGAAAGCAUCAAUUCCUUUUCGGCCGCAUCUGACCCGAUCUUAGCUGCAACAAUCUGU ..(((((.(((((((((((..........))))))................(((((.(((..((((((.(.....).)))))).))))))))..........))))).)))))....... (-25.80 = -26.44 + 0.64)

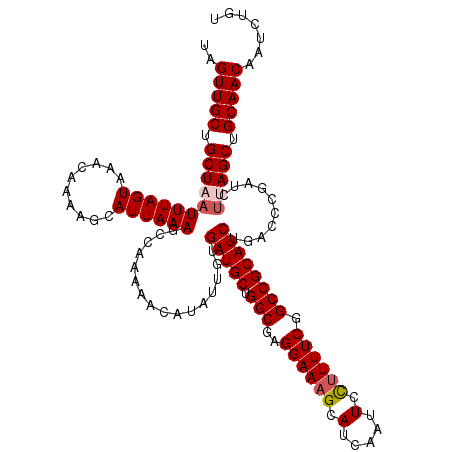
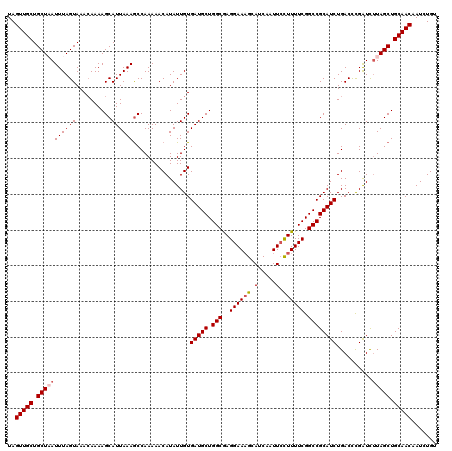
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:54:37 2006