
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,716,651 – 2,716,748 |
| Length | 97 |
| Max. P | 0.761932 |

| Location | 2,716,651 – 2,716,748 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 83.37 |
| Mean single sequence MFE | -18.71 |
| Consensus MFE | -16.28 |
| Energy contribution | -16.33 |
| Covariance contribution | 0.06 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.56 |
| Structure conservation index | 0.87 |
| SVM decision value | 0.35 |
| SVM RNA-class probability | 0.701833 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2716651 97 + 22407834 AU------ACUCAAUAUUCAGUUGAGCAAAUCCAUGCGUUGUUCGGUUGCAAUUCGCAUGCAUUCACUCGGUUCCUUUUUUCGCUUGUUUUAUUUCCA------------UU--UUU--- ..------.............(((((..(((.((((((((((......))))..)))))).)))..)))))...........................------------..--...--- ( -14.90) >DroGri_CAF1 172234 110 + 1 AUAGUCGCACUCAAUAUUCAGUUGAGCAAAUCCAUCCGUUGUUCUCUUGCAAUUCGCAUGCAUUCACUUGGCUCGUUUCG--GAUUGCUUC--UGCCCC-A-----CUUGCCACUUGCCA ......((.((((((.....))))))...........(((((......)))))..))..(((......((((..((..((--((.....))--))....-)-----)..))))..))).. ( -19.20) >DroSec_CAF1 135533 105 + 1 AU------ACUCAAUAUUCAGGUGAGCAAAUCCAUGCGUUGUUCGGUUGCAAUUCGCAUGCAUUCACUCGGCUCCUUUUUUCGCUUGUUUUAUUUCCCC-A-----UUUCUU--UUUUU- ..------....((((..((((((((..(((.((((((((((......))))..)))))).)))..)))(....).......)))))...)))).....-.-----......--.....- ( -18.00) >DroSim_CAF1 139049 105 + 1 AU------ACUCAAUAUUCAGUUGAGCAAAUCCAUGCGUUGUUCGGUUGCAAUUCGCAUGCAUUCACUCGGCUCCUUUUUUCGCUUGUUUUAUUUCCCC-A-----UUUCUU--UUUUU- ..------...........(((((((..(((.((((((((((......))))..)))))).)))..)))))))..........................-.-----......--.....- ( -19.10) >DroEre_CAF1 153778 109 + 1 AU------ACUCAAUAUUCAGUUGAGCAAAUCCAUGCGUUGUUCGCUUGCAAUUCGCAUGCAUUCACUCGGCUCCUUUUUUCGCUUGUUUUAUUUCCCCCACAUCUAUUUUU--UUU--- ..------...........(((((((..(((.((((((((((......))))..)))))).)))..))))))).......................................--...--- ( -19.10) >DroYak_CAF1 143186 111 + 1 AU------ACUCAAUAUUCAGUUGAGCAAAUCCAUGCGUUGUUCGGUUGCAAUUCGCAUGCAUUCACUCGGCUCCUUUUUUCGCUUGUUUUAUUUUCCCCAAAGCAUUUUUU--UUUUG- ..------...........(((((((..(((.((((((((((......))))..)))))).)))..))))))).........((((...............)))).......--.....- ( -21.96) >consensus AU______ACUCAAUAUUCAGUUGAGCAAAUCCAUGCGUUGUUCGGUUGCAAUUCGCAUGCAUUCACUCGGCUCCUUUUUUCGCUUGUUUUAUUUCCCC_A_____UUUCUU__UUUU__ ...................(((((((..(((.((((((((((......))))..)))))).)))..)))))))............................................... (-16.28 = -16.33 + 0.06)
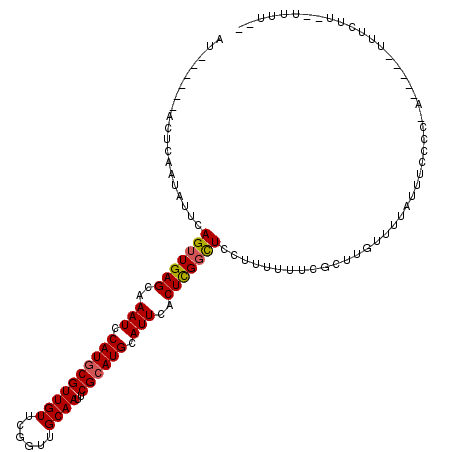


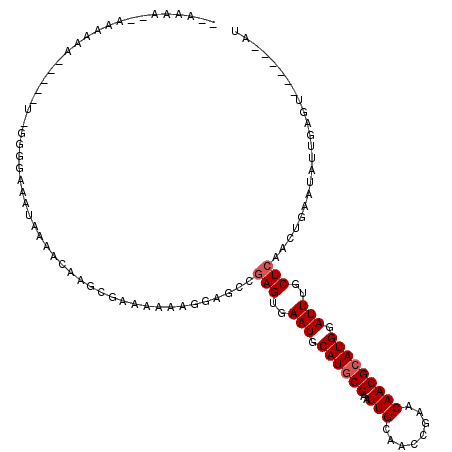
| Location | 2,716,651 – 2,716,748 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 83.37 |
| Mean single sequence MFE | -22.53 |
| Consensus MFE | -17.63 |
| Energy contribution | -17.97 |
| Covariance contribution | 0.33 |
| Combinations/Pair | 1.00 |
| Mean z-score | -2.01 |
| Structure conservation index | 0.78 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.761932 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2716651 97 - 22407834 ---AAA--AA------------UGGAAAUAAAACAAGCGAAAAAAGGAACCGAGUGAAUGCAUGCGAAUUGCAACCGAACAACGCAUGGAUUUGCUCAACUGAAUAUUGAGU------AU ---...--..------------.............................(((..(((.((((((..(((........))))))))).)))..)))...............------.. ( -19.70) >DroGri_CAF1 172234 110 - 1 UGGCAAGUGGCAAG-----U-GGGGCA--GAAGCAAUC--CGAAACGAGCCAAGUGAAUGCAUGCGAAUUGCAAGAGAACAACGGAUGGAUUUGCUCAACUGAAUAUUGAGUGCGACUAU .....(((.(((((-----(-.(((((--((....(((--((..((.......))...((((.......)))).........)))))...))))))).)))..........))).))).. ( -25.20) >DroSec_CAF1 135533 105 - 1 -AAAAA--AAGAAA-----U-GGGGAAAUAAAACAAGCGAAAAAAGGAGCCGAGUGAAUGCAUGCGAAUUGCAACCGAACAACGCAUGGAUUUGCUCACCUGAAUAUUGAGU------AU -.....--......-----(-.(((...........((..........)).(((..(((.((((((..(((........))))))))).)))..))).))).).........------.. ( -23.20) >DroSim_CAF1 139049 105 - 1 -AAAAA--AAGAAA-----U-GGGGAAAUAAAACAAGCGAAAAAAGGAGCCGAGUGAAUGCAUGCGAAUUGCAACCGAACAACGCAUGGAUUUGCUCAACUGAAUAUUGAGU------AU -.....--......-----.-............(((((..........)).(((..(((.((((((..(((........))))))))).)))..))).........)))...------.. ( -20.40) >DroEre_CAF1 153778 109 - 1 ---AAA--AAAAAUAGAUGUGGGGGAAAUAAAACAAGCGAAAAAAGGAGCCGAGUGAAUGCAUGCGAAUUGCAAGCGAACAACGCAUGGAUUUGCUCAACUGAAUAUUGAGU------AU ---...--.......(((((..((............((..........)).(((..(((.((((((..(((....)))....)))))).)))..)))..))..)))))....------.. ( -23.90) >DroYak_CAF1 143186 111 - 1 -CAAAA--AAAAAAUGCUUUGGGGAAAAUAAAACAAGCGAAAAAAGGAGCCGAGUGAAUGCAUGCGAAUUGCAACCGAACAACGCAUGGAUUUGCUCAACUGAAUAUUGAGU------AU -.....--.....((((((.................((..........)).(((..(((.((((((..(((........))))))))).)))..)))...........))))------)) ( -22.80) >consensus __AAAA__AAAAAA_____U_GGGGAAAUAAAACAAGCGAAAAAAGGAGCCGAGUGAAUGCAUGCGAAUUGCAACCGAACAACGCAUGGAUUUGCUCAACUGAAUAUUGAGU______AU ...................................................(((..(((.((((((..(((........))))))))).)))..)))....................... (-17.63 = -17.97 + 0.33)


Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:49:18 2006