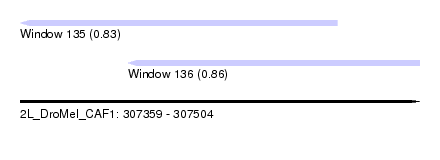
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 307,359 – 307,504 |
| Length | 145 |
| Max. P | 0.864977 |

| Location | 307,359 – 307,474 |
|---|---|
| Length | 115 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 82.53 |
| Mean single sequence MFE | -44.67 |
| Consensus MFE | -25.91 |
| Energy contribution | -30.48 |
| Covariance contribution | 4.56 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.46 |
| Structure conservation index | 0.58 |
| SVM decision value | 0.72 |
| SVM RNA-class probability | 0.833493 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 307359 115 - 22407834 AGAAUGAGCAAGAGGGGGGGGGGGUUUCC----AGGGGUUUGGCCCCCACUGCUUUCCCGCUCUCUCCCUCAUCACAAUGACGCGUGGUUAUUGUAAUUAAAGUUAUGUGCGAUAUGUG- .((.((((..((((((.(((..(((....----.((((.....))))....)))..))).))))))..))))))((((((((.....))))))))........................- ( -44.30) >DroSec_CAF1 11653 112 - 1 AGAAAGAACGAGAGGGGGGGCG---UUUC----AGGGGUUGGGCCCCCACUGCCUUCCCUCUCUGUCGCUCAUCACGAUGACGCGUGGUUAUUGUAAUUAAAGUUAUGUGCGAUAUGUG- .........(((((((((((((---....----.((((.....))))...)))))))))))))((((((.(((.((((((((.....))))))))(((....)))))).))))))....- ( -46.00) >DroEre_CAF1 12241 108 - 1 -GCGAGAGCG------AGGCAGGGUGGUC----AGGGGUGGGGCC-CCACUGCUUUCCCUCUCUGUUCCUCAUCACGAUGACGCGUGGUUAUUGUAAUUAAAGUUAUGUGCGAUAUGUGC -(((..((((------((((((((.((..----((.(((((....-))))).))..))..)))))..))))...((((((((.....)))))))).......)))...)))......... ( -38.30) >DroYak_CAF1 12490 118 - 1 -GCGAGAACGGGAGGGAAGGAGGGUGGUUGGAAAGGGGUGGGGCCCCCACUGCUUUCCCUCUCUGUCCCUCAUCACGAUGACGCGUGGUUAUUGUAAUUAAAGUUAUGUGCGAUAUGUG- -(((..(((((((((((.((((((.....((((((.((((((...)))))).)))))))))))).)))))).))((((((((.....)))))))).......)))...)))........- ( -50.10) >consensus _GAAAGAACGAGAGGGAGGGAGGGUGGUC____AGGGGUGGGGCCCCCACUGCUUUCCCUCUCUGUCCCUCAUCACGAUGACGCGUGGUUAUUGUAAUUAAAGUUAUGUGCGAUAUGUG_ .....(((.((.((((((((((((((((......((((.....)))).)))))))))))))))).)).)))(((((((((((.....)))))))).......((.....)))))...... (-25.91 = -30.48 + 4.56)

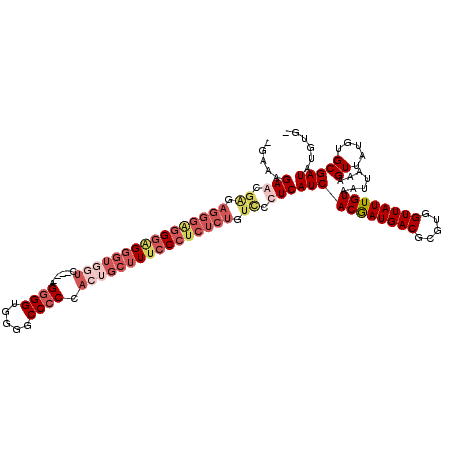
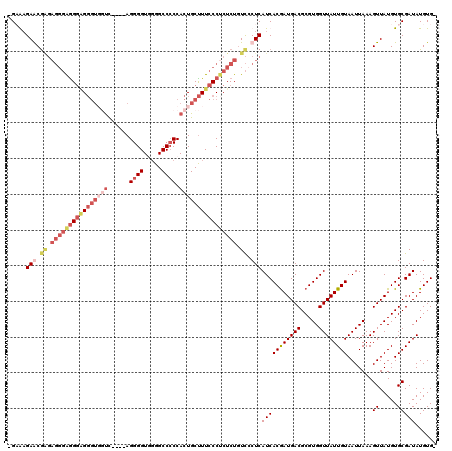
| Location | 307,398 – 307,504 |
|---|---|
| Length | 106 |
| Sequences | 4 |
| Columns | 114 |
| Reading direction | reverse |
| Mean pairwise identity | 75.78 |
| Mean single sequence MFE | -46.80 |
| Consensus MFE | -22.65 |
| Energy contribution | -27.90 |
| Covariance contribution | 5.25 |
| Combinations/Pair | 1.13 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.48 |
| SVM decision value | 0.84 |
| SVM RNA-class probability | 0.864977 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
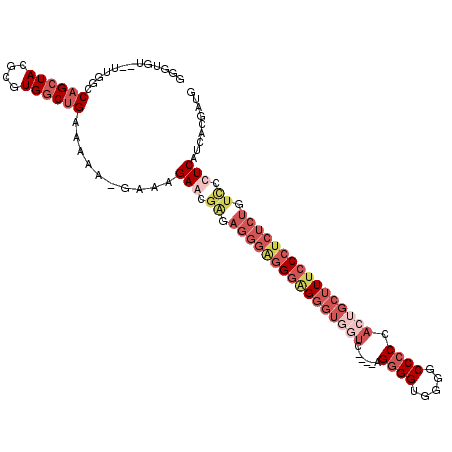
>2L_DroMel_CAF1 307398 106 - 22407834 GUAUGUUUU----CAGCUACGCGUGGCUGAAAAAAGAAUGAGCAAGAGGGGGGGGGGGUUUCC----AGGGGUUUGGCCCCCACUGCUUUCCCGCUCUCUCCCUCAUCACAAUG .....((((----((((((....))))))))))..((.((((..((((((.(((..(((....----.((((.....))))....)))..))).))))))..))))))...... ( -49.20) >DroSec_CAF1 11692 107 - 1 GGGUGUUUUCGGCCAGCUACGCGUGGCUGAAAAAAGAAAGAACGAGAGGGGGGGCG---UUUC----AGGGGUUGGGCCCCCACUGCCUUCCCUCUCUGUCGCUCAUCACGAUG (((((((((((((((((...)).))))))))))..........(((((((((((((---....----.((((.....))))...)))))))))))))...)))))......... ( -51.00) >DroEre_CAF1 12281 99 - 1 GGGU-U--UUGGCCAGCUACGCGUGCCUGAAAAU-GCGAGAGCG------AGGCAGGGUGGUC----AGGGGUGGGGCC-CCACUGCUUUCCCUCUCUGUUCCUCAUCACGAUG .(((-.--...))).(((.(((((........))-)))..)))(------((((((((.((..----((.(((((....-))))).))..))..)))))..))))......... ( -37.80) >DroYak_CAF1 12529 111 - 1 GGUUGU--UUGGCCUGCUACGCGUGGCUGAAAAA-GCGAGAACGGGAGGGAAGGAGGGUGGUUGGAAAGGGGUGGGGCCCCCACUGCUUUCCCUCUCUGUCCCUCAUCACGAUG (.((.(--(..((((((...))).)))..)).))-.).....((.((((((.((((((.....((((((.((((((...)))))).)))))))))))).))))))....))... ( -49.20) >consensus GGGUGU__UUGGCCAGCUACGCGUGGCUGAAAAA_GAAAGAACGAGAGGGAGGGAGGGUGGUC____AGGGGUGGGGCCCCCACUGCUUUCCCUCUCUGUCCCUCAUCACGAUG .............((((((....))))))..........(((.((.((((((((((((((((......((((.....)))).)))))))))))))))).)).)))......... (-22.65 = -27.90 + 5.25)


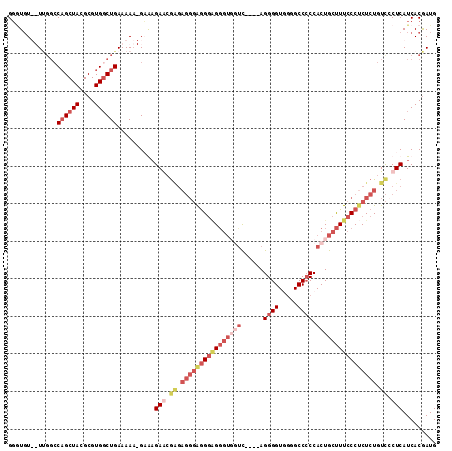
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:25:29 2006