
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,469,322 – 2,469,426 |
| Length | 104 |
| Max. P | 0.880795 |

| Location | 2,469,322 – 2,469,426 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 111 |
| Reading direction | reverse |
| Mean pairwise identity | 87.55 |
| Mean single sequence MFE | -33.35 |
| Consensus MFE | -28.00 |
| Energy contribution | -28.00 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.11 |
| Mean z-score | -2.33 |
| Structure conservation index | 0.84 |
| SVM decision value | 0.91 |
| SVM RNA-class probability | 0.880795 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
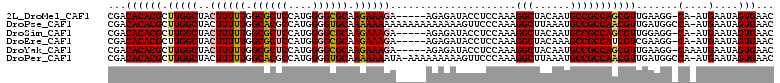
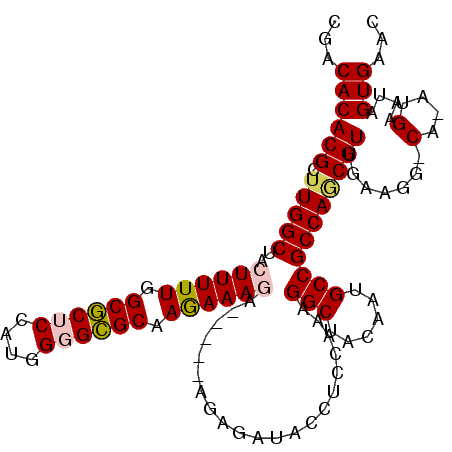
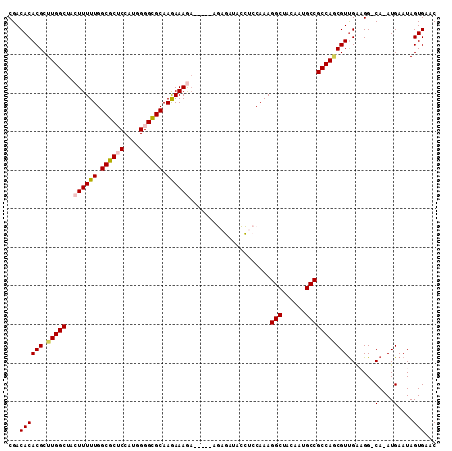
>2L_DroMel_CAF1 2469322 104 - 22407834 CGACACACGCUUGGCUACUUUUUGGCGCUCCAUGGGGCGCAAGAAAGA-----AGAGAUACCUCCAAAGGCUACAAUGCCGCCAGCGUUGAAGG-CA-AUGAAUAGUGAAC ...((((((((.(((..((((((.((((((....)))))).)))))).-----.(((....)))....(((......)))))))))))((....-))-.......)))... ( -36.00) >DroPse_CAF1 55981 110 - 1 CGACACACGCUUGGCUACUUUUUGGCACGCCAUGGGGUGCAAAAAAAAAAAAAAAAAAAGUUCCCAAAGGCUUAAAUGCCGCCAACGUUGAUGGCCA-AUGAAUAGUGAAC .....(((((((((..(((((((.((((.(....).))))..............)))))))..)))).(((......)))))...(((((.....))-)))....)))... ( -28.96) >DroSim_CAF1 50101 104 - 1 CGACACACGCUUGGCUACUUUUUGGCGCUCCAUGGGGCGCAAGAAAGA-----AGAGAUACCUCCAAAGGCUACAAUGCCGCCAGCGUUGGAGG-CA-AUGAAUAGUGAAC (..((.(((((.(((..((((((.((((((....)))))).)))))).-----.(((....)))....(((......)))))))))))))..).-..-............. ( -36.80) >DroEre_CAF1 50694 104 - 1 CGACACACGCUUGGCUACUUUUUGGCGCUCCAUGGGGCGCAAGAAAGA-----AGAGAUACCUCCAAAGGCUACAAAGCCGCCAUCGUCGAAGG-CA-AUGAAUAGUGAAC ...((((((..((((..((((((.((((((....)))))).)))))).-----.(((....)))....((((....)))))))).)))(....)-..-.......)))... ( -33.30) >DroYak_CAF1 50872 105 - 1 CGACACACGCUUGGCUACUUUUUGGCGCUCCAUGGGGCGCAAGAAAGA-----AGAGAUACCUCCAAAGGCUACAAUGCCGCCAGCGUUGAAGG-CAAAUGAAUAGUGAAC ...((((((((.(((..((((((.((((((....)))))).)))))).-----.(((....)))....(((......)))))))))))((....-))........)))... ( -36.00) >DroPer_CAF1 54487 109 - 1 CGACACACGCUUGGCUACUUUUUGGCACGCCAUGGGGUGCAAAAAAAUA-AAAAAAAAAGUUCCCAAAGGCUUAAAUGCCGCCAACGUUGAUGGCCA-AUGAAUAGUGAAC .....(((((((((..(((((((.((((.(....).)))).........-....)))))))..)))).(((......)))))...(((((.....))-)))....)))... ( -29.04) >consensus CGACACACGCUUGGCUACUUUUUGGCGCUCCAUGGGGCGCAAGAAAGA_____AGAGAUACCUCCAAAGGCUACAAUGCCGCCAGCGUUGAAGG_CA_AUGAAUAGUGAAC ...((((((.(((((..((((((.((((((....)))))).)))))).....................(((......))))))))))).......(....)....)))... (-28.00 = -28.00 + 0.00)
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:47:31 2006