| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,348,804 – 2,348,908 |
| Length | 104 |
| Max. P | 0.900491 |

| Location | 2,348,804 – 2,348,908 |
|---|---|
| Length | 104 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 78.35 |
| Mean single sequence MFE | -26.58 |
| Consensus MFE | -16.61 |
| Energy contribution | -16.08 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.98 |
| Structure conservation index | 0.62 |
| SVM decision value | 1.01 |
| SVM RNA-class probability | 0.900491 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 2348804 104 - 22407834 ---------UUGUU-CCUUC--UGCUCGGGCGAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACGAACAUAAAUUUGUUGCUCGUAAAUCCACUCGC--CAGACCAAAACACAAAAA ---------((((.-..((.--((.((.(((((....((.(....)))........(..(((((((((......)))))))))..)........))))--).)).)).)).))))... ( -24.70) >DroVir_CAF1 93169 103 - 1 GGGUAUGAAGUGUC-CUGGC--GG-CAUGGCCAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACAAACAUAAAUUUGUUGCUCGUAAAUCCACUGGC---GGACC----CAGU---- ((((.((.((((..-.((((--..-....))))....((.(....)))........(..(((((((((......)))))))))..).....))))..)---).)))----)...---- ( -30.80) >DroPse_CAF1 110084 103 - 1 ---------GUGCUACCCAG--UGCUCGGGCGAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACGUACAUAAAUUUGUUGCUCGUAAAUCCACUUGCCCCAGACC----CAGAAAAA ---------(..((....))--..)..((((((....((.(....)))........(..(((((((..........)))))))..)........))))))......----........ ( -24.00) >DroGri_CAF1 63662 104 - 1 GGUAGUGAAGUGUU-CUGUU--GGCCAUGGCCAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACAAACAUAAAUUUGUUGCUCGUAAAUCCACUGGC---GGACC----CAGU---- (.(((((..((((.-((..(--(((....))))...)).)))).............(..(((((((((......)))))))))..).....))))).)---.....----....---- ( -30.00) >DroAna_CAF1 59232 105 - 1 ----------GGCU-CGGACGGUGCUCGGACGAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACGAACAUAAAUUUGUUGCUCGUAAAUCCACUUGC--CAGACCAAAACACAAAAG ----------((((-.((..((((.....((((....((.(....)))............((((((((......)))))))))))).....))))..)--))).))............ ( -26.00) >DroPer_CAF1 121036 103 - 1 ---------GUGCUACCCAG--UGCUCGGGCGAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACGUACAUAAAUUUGUUGCUCGUAAAUCCACUUGCCCCAGACC----CAGAAAAA ---------(..((....))--..)..((((((....((.(....)))........(..(((((((..........)))))))..)........))))))......----........ ( -24.00) >consensus _________GUGCU_CCGAC__UGCUCGGGCGAUUAAGCACGCAAGGCAAAUAAAAAUAGGCAACGAACAUAAAUUUGUUGCUCGUAAAUCCACUUGC__CAGACC____CAGAAAAA ........................((...((((....((.(....)))........(..(((((((((......)))))))))..)........))))...))............... (-16.61 = -16.08 + -0.53)

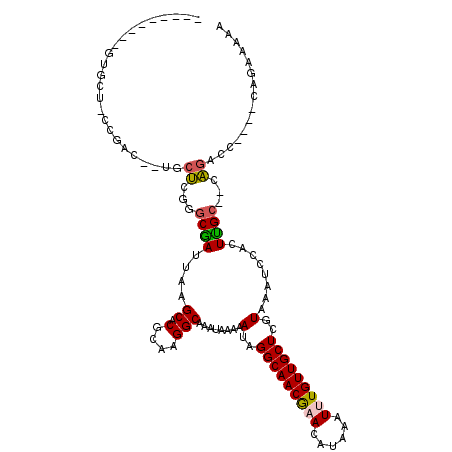
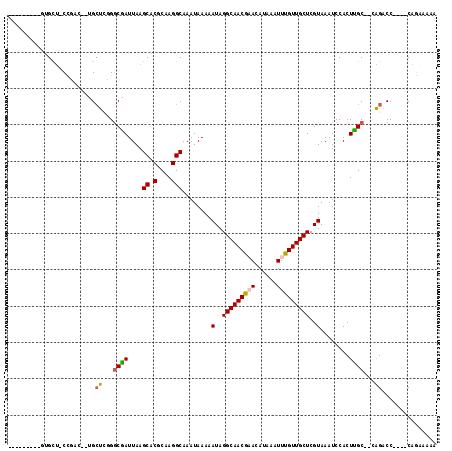
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:46:26 2006