
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 282,033 – 282,131 |
| Length | 98 |
| Max. P | 0.500000 |

| Location | 282,033 – 282,131 |
|---|---|
| Length | 98 |
| Sequences | 4 |
| Columns | 99 |
| Reading direction | forward |
| Mean pairwise identity | 91.33 |
| Mean single sequence MFE | -17.13 |
| Consensus MFE | -13.06 |
| Energy contribution | -13.57 |
| Covariance contribution | 0.50 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.76 |
| SVM decision value | -0.06 |
| SVM RNA-class probability | 0.500000 |
| Prediction | OTHER |
Download alignment: ClustalW | MAF
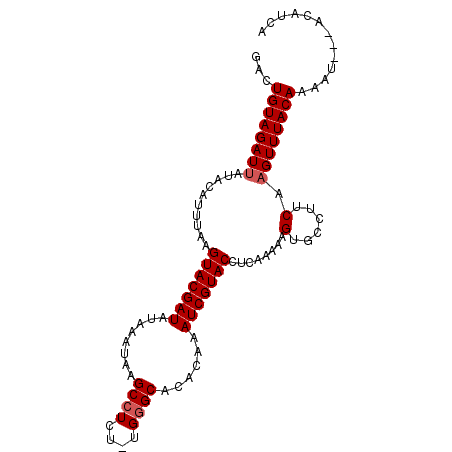
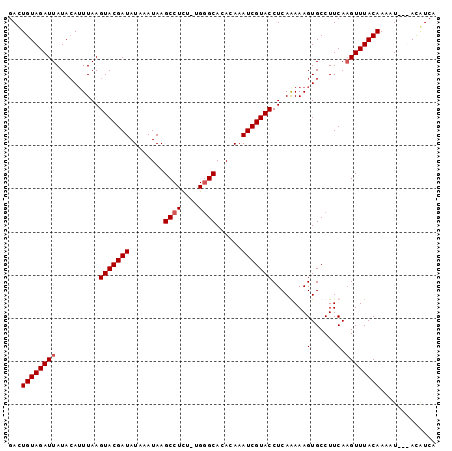
>2L_DroMel_CAF1 282033 98 + 22407834 GAUUGUAGAUUAUAGAUUUAAGUACGAUAUAAAUAAGCCUCU-UGGGCACACAAAUCGUACCUCAGAAAGUGCCUUCAAGUUUACAAAAUUAUACAUAA ..(((((((((..........(((((((........(((...-..)))......)))))))....(((......))).)))))))))............ ( -18.14) >DroSec_CAF1 4092 99 + 1 GAAUGUAGAUUAUACAUUUAAGUACGAUAUAAAUAAGCCUCUUUGGGCACACAAAUCGUACCUCAGAAAGUGCCUUCAAGUUUACAAAAUCAUACAUCA (((((((.....)))))))..(((.(((.....(((((......((((((.....((........))..))))))....)))))....))).))).... ( -17.60) >DroEre_CAF1 4162 93 + 1 GACUGUAGAUUAUACAUUUAAGUACGAUAUAAAUAAGCCUCU-UG-GCACACAAAUCGUACCUCAAAAAGUGCCUUCA-GUUUACAAAAU---AUGUCA (((((((((((...(((((..(((((((........(((...-.)-))......)))))))......))))).....)-)))))))....---..))). ( -17.04) >DroYak_CAF1 4128 95 + 1 GACUGUAGAUUAUACAAUUAAGUACGAUAUAAAUAAGCCUUU-UGGGCACACAAAUCGUACCUCAAAAAGUGCCUUCAAGUUUACAAAAU---AUGUCA (((((((((((..........(((((((........(((...-..)))......)))))))........(......).))))))))....---..))). ( -15.74) >consensus GACUGUAGAUUAUACAUUUAAGUACGAUAUAAAUAAGCCUCU_UGGGCACACAAAUCGUACCUCAAAAAGUGCCUUCAAGUUUACAAAAU___ACAUCA ...((((((((..........(((((((........((((....))))......)))))))........(......).))))))))............. (-13.06 = -13.57 + 0.50)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:25:15 2006