
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 2,247,237 – 2,247,340 |
| Length | 103 |
| Max. P | 0.805596 |

| Location | 2,247,237 – 2,247,340 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | forward |
| Mean pairwise identity | 79.01 |
| Mean single sequence MFE | -27.18 |
| Consensus MFE | -17.25 |
| Energy contribution | -16.67 |
| Covariance contribution | -0.58 |
| Combinations/Pair | 1.28 |
| Mean z-score | -1.70 |
| Structure conservation index | 0.63 |
| SVM decision value | 0.55 |
| SVM RNA-class probability | 0.778334 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
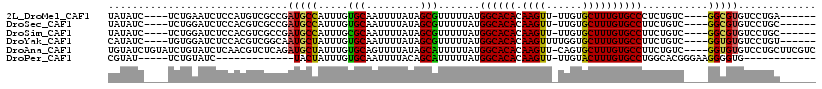
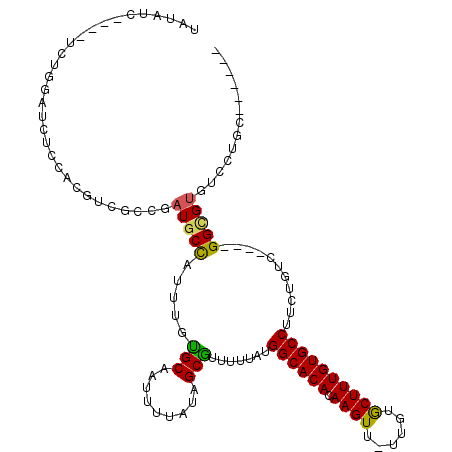
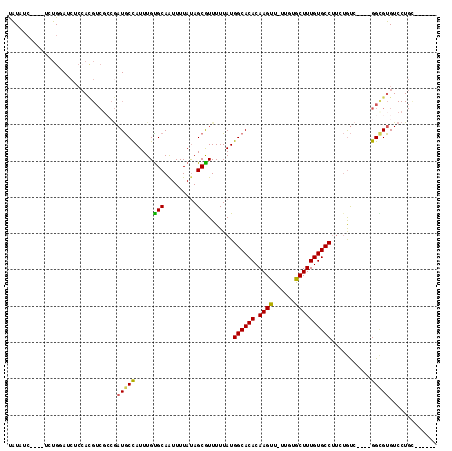
>2L_DroMel_CAF1 2247237 103 + 22407834 UAUAUC----UCUGAAUCUCCAUGUCGCCGAUGCCAUUUGUGCAAUUUUAUAGCGUUUUUAUGGCACACAAGUU-UUGUGCUUUGUGCCCUCUGUC----GGCGUGUCCUGA------ ......----..........((.(.((((((((((....).))...................((((((.((((.-....))))))))))....)))----))))...).)).------ ( -24.70) >DroSec_CAF1 7533 103 + 1 UAUAUC----UCUGGAUCUCCACGUCGCCGAUGCCAUUUGUGCAAUUUUAUAGCGUUUUUAUGGCACACAAGUU-UUGUGCUUUGUGCCUUCUGUC----GGCGUGUCCUGC------ ......----...((....))(((.((((((((((....).))...................((((((.((((.-....))))))))))....)))----))))))).....------ ( -27.70) >DroSim_CAF1 7781 103 + 1 UAUAUC----UCUGGAUCUCCACGUCGCCGAUGCCAUUUGCGCAAUUUUAUAGCGUUUUUAUGGCACACAAGUU-UUGUGCUUUGUGCCUUCUGUC----GGCGUGUCCUGC------ ......----...((....))(((.((((((((((((..((((.........))))....))))).(((((((.-....)).)))))......)))----))))))).....------ ( -31.00) >DroYak_CAF1 7823 104 + 1 CAUAUC----UGUGGAUCUCCACGUCGGCAAUGCUAUUUGUGCAAUUUUAUAGCGUUUUUAUGGCACACAAGUUUUGGUGCUUUGUGCCUUCUGUC----GGUGUGUCCUGU------ ((((((----.((((....))))...(((((((((((..(......)..)))))))).....((((((.((((......))))))))))....)))----))))))......------ ( -32.20) >DroAna_CAF1 8562 113 + 1 UGUAUCUGUAUCUGUAUCUCAACGUCUCAGAUGCUAUUUGUGCAGUUUUAUAGCAUUUUUAUGGCACACAAGUU-CAGUGCUUUGUGCCUUCUGUC----GGUGUGUCCUGCUUCGUC .(((((((....(((......)))...))))))).......((((...((((.(........((((((.((((.-....)))))))))).......----).))))..))))...... ( -27.46) >DroPer_CAF1 7119 87 + 1 CGUAU-----UCUGUAUC-------------UACUAUUUGUGCAAUUUUACAGCAUUUUUAUGGCACACAAGUU-UUGUACUUUGUGCCUGGCACGGGAAGGGGUG------------ .....-----....((((-------------(.((.(((((((...................((((((.((((.-....))))))))))..))))))).)))))))------------ ( -20.00) >consensus UAUAUC____UCUGGAUCUCCACGUCGCCGAUGCCAUUUGUGCAAUUUUAUAGCGUUUUUAUGGCACACAAGUU_UUGUGCUUUGUGCCUUCUGUC____GGCGUGUCCUGC______ ..............................(((((.....(((.........))).......((((((.((((......))))))))))...........)))))............. (-17.25 = -16.67 + -0.58)
| Location | 2,247,237 – 2,247,340 |
|---|---|
| Length | 103 |
| Sequences | 6 |
| Columns | 118 |
| Reading direction | reverse |
| Mean pairwise identity | 79.01 |
| Mean single sequence MFE | -23.85 |
| Consensus MFE | -14.31 |
| Energy contribution | -13.78 |
| Covariance contribution | -0.53 |
| Combinations/Pair | 1.21 |
| Mean z-score | -1.90 |
| Structure conservation index | 0.60 |
| SVM decision value | 0.63 |
| SVM RNA-class probability | 0.805596 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
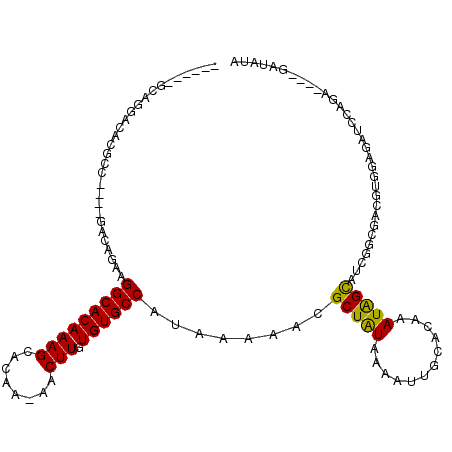
>2L_DroMel_CAF1 2247237 103 - 22407834 ------UCAGGACACGCC----GACAGAGGGCACAAAGCACAA-AACUUGUGUGCCAUAAAAACGCUAUAAAAUUGCACAAAUGGCAUCGGCGACAUGGAGAUUCAGA----GAUAUA ------........((((----((.....(((((((((.....-..))).))))))........(((((............))))).))))))...............----...... ( -24.70) >DroSec_CAF1 7533 103 - 1 ------GCAGGACACGCC----GACAGAAGGCACAAAGCACAA-AACUUGUGUGCCAUAAAAACGCUAUAAAAUUGCACAAAUGGCAUCGGCGACGUGGAGAUCCAGA----GAUAUA ------........((((----((.....(((((((((.....-..))).))))))........(((((............))))).))))))...(((....)))..----...... ( -27.30) >DroSim_CAF1 7781 103 - 1 ------GCAGGACACGCC----GACAGAAGGCACAAAGCACAA-AACUUGUGUGCCAUAAAAACGCUAUAAAAUUGCGCAAAUGGCAUCGGCGACGUGGAGAUCCAGA----GAUAUA ------........((((----((.....(((((((((.....-..))).))))))........(((((............))))).))))))...(((....)))..----...... ( -27.30) >DroYak_CAF1 7823 104 - 1 ------ACAGGACACACC----GACAGAAGGCACAAAGCACCAAAACUUGUGUGCCAUAAAAACGCUAUAAAAUUGCACAAAUAGCAUUGCCGACGUGGAGAUCCACA----GAUAUG ------...((.((....----.......(((((((((........))).))))))........(((((............)))))..))))...((((....)))).----...... ( -22.90) >DroAna_CAF1 8562 113 - 1 GACGAAGCAGGACACACC----GACAGAAGGCACAAAGCACUG-AACUUGUGUGCCAUAAAAAUGCUAUAAAACUGCACAAAUAGCAUCUGAGACGUUGAGAUACAGAUACAGAUACA ((((.....((.....))----..((((.(((((((((.....-..))).)))))).......((((((............))))))))))...)))).................... ( -22.50) >DroPer_CAF1 7119 87 - 1 ------------CACCCCUUCCCGUGCCAGGCACAAAGUACAA-AACUUGUGUGCCAUAAAAAUGCUGUAAAAUUGCACAAAUAGUA-------------GAUACAGA-----AUACG ------------..........((((...((((((((((....-.)))).)))))).........(((((...((((.......)))-------------).))))).-----.)))) ( -18.40) >consensus ______GCAGGACACGCC____GACAGAAGGCACAAAGCACAA_AACUUGUGUGCCAUAAAAACGCUAUAAAAUUGCACAAAUAGCAUCGGCGACGUGGAGAUCCAGA____GAUAUA .............................(((((((((........))).))))))........(((((............)))))................................ (-14.31 = -13.78 + -0.53)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 09:45:33 2006