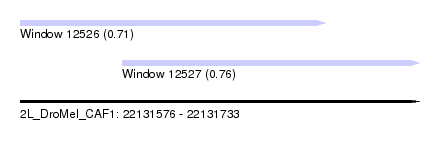

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 22,131,576 – 22,131,733 |
| Length | 157 |
| Max. P | 0.761857 |

| Location | 22,131,576 – 22,131,696 |
|---|---|
| Length | 120 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 80.83 |
| Mean single sequence MFE | -44.70 |
| Consensus MFE | -30.05 |
| Energy contribution | -29.61 |
| Covariance contribution | -0.44 |
| Combinations/Pair | 1.36 |
| Mean z-score | -1.65 |
| Structure conservation index | 0.67 |
| SVM decision value | 0.36 |
| SVM RNA-class probability | 0.706189 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
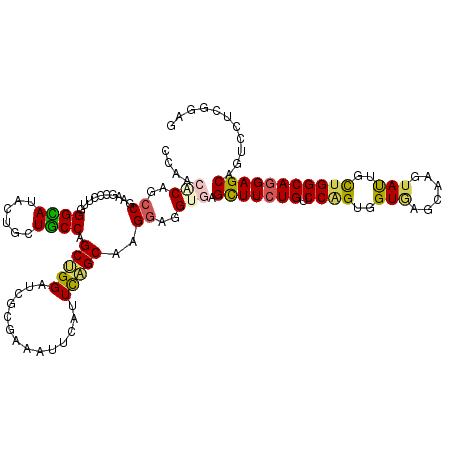
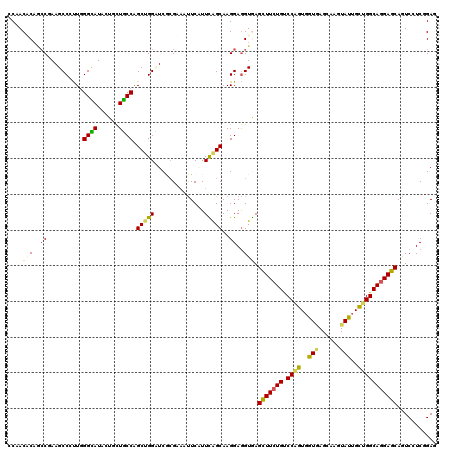
>2L_DroMel_CAF1 22131576 120 + 22407834 ACAACGCUCCCGAAGCCGCUGGGGAUUCUGCUUCCAGCCGGAUCGCGCCAUUCUUUUGGCAAGGAGGUAAGCUUCUGCCCAGUGGUAAGCAAGUACUGUUGGCAGGAGCAGUCCUCCGAG .....((.((((.......))))(((((.((.....)).)))))))((((......))))..(((((...(((((((((.((..(((......)))..)))))))))))...)))))... ( -51.20) >DroVir_CAF1 4152 120 + 1 CCGACACAGCCAAAGCCCUUGGGCAUACUGUUGCCGGCGGGCUCGCGUAAUUCAUUCAGCAAGGAGGUGAGUUUCUGUCCAGUGGUGAGUAAGUAUUGCUGGCAGGAGCAAUCGUCGGAG (((((((..((..(((((.(.((((......)))).).))))).((............))..))..))..(((((((.((((..(((......)))..)))))))))))....))))).. ( -50.40) >DroGri_CAF1 4761 120 + 1 GCAACUCAAGCGAAGCCUUUGGGCAUACUGCUGCCAGCUGGUUCGCGCAAUUCGUUCAGCAAGGAGGUGAGUUUCUGUCCGGUGGUGAGCAAGUAUUGCUGGCAGGAGCAGUCAUCGGAG ....(((..(((((.((....((((......))))....)))))))...................((((((((((((.((((..(((......)))..)))))))))))..))))).))) ( -41.70) >DroWil_CAF1 2481 120 + 1 CCCACUCAGCCCAAGCCCUUGGGUAUUCUCCUGCCAGCUGGAUCGAGAAAUUCCUUCAGCAAGGAGGUGAGUUUCUGUCCAGUCGUUAGCAAGUAUUGCUGGCAAGAGCAAUCCUCUGAA .....(((((((((....)))))....(((.((((.(((((((..((((((((((((......)))).)))))))))))))))....((((.....)))))))).))).......)))). ( -42.60) >DroMoj_CAF1 2578 120 + 1 CCGACACAGCCGAAGCCACUGGGCAUACUCCUGCCGGCUGGCUCACGUAAUUCGUUUAGCAAGGAGGUGAGCUUCUGUCCCGUGGUGAGUAAGUAUUGCUGGCAGGAGCAGUCGUCGGAG (((((((.(((.(......).)))...(((((((((((..((((((.......(.....)..(((((....)))))........)))))).......)))))))))))..)).))))).. ( -47.90) >DroAna_CAF1 21111 120 + 1 GCCACGCAACCCAAGCCUUUAGGCAUCCUACUGCCAGCUGGUUCCCGAAACUCAUUUAGCAAGGAAGUAAGCUUCUGUCCAGUGGUAAGCAAGUACUGUUGGCAGGAGCAGUCCUCCGAA .....((......((((....((((......))))....))))...(.....).....))..(((.(...(((((((.((((..(((......)))..)))))))))))...).)))... ( -34.40) >consensus CCAACACAGCCGAAGCCCUUGGGCAUACUGCUGCCAGCUGGAUCGCGAAAUUCAUUCAGCAAGGAGGUGAGCUUCUGUCCAGUGGUGAGCAAGUAUUGCUGGCAGGAGCAGUCCUCGGAG ....(((..((..........((((......)))).(((((..............)))))..))..))).(((((((.((((..(((......)))..)))))))))))........... (-30.05 = -29.61 + -0.44)
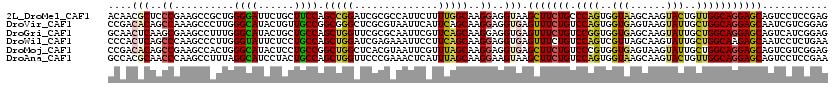

| Location | 22,131,616 – 22,131,733 |
|---|---|
| Length | 117 |
| Sequences | 6 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 77.55 |
| Mean single sequence MFE | -41.82 |
| Consensus MFE | -24.70 |
| Energy contribution | -23.57 |
| Covariance contribution | -1.13 |
| Combinations/Pair | 1.53 |
| Mean z-score | -1.71 |
| Structure conservation index | 0.59 |
| SVM decision value | 0.50 |
| SVM RNA-class probability | 0.761857 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
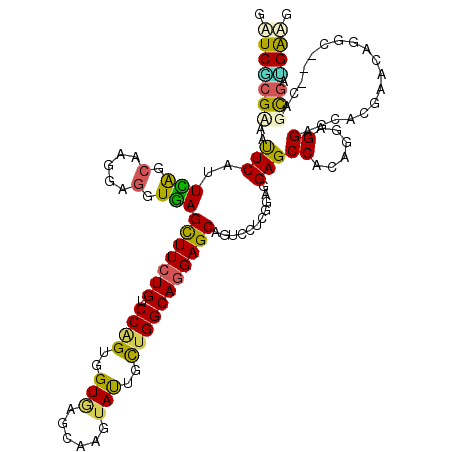
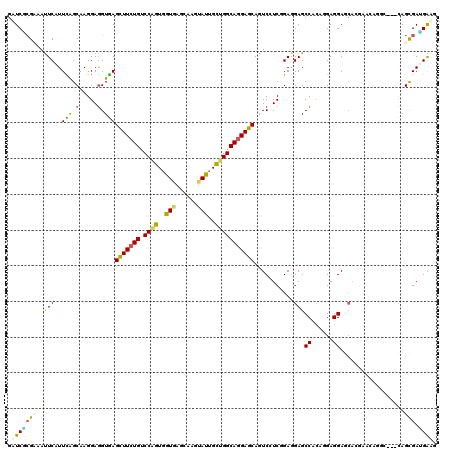
>2L_DroMel_CAF1 22131616 117 + 22407834 GAUCGCGCCAUUCUUUUGGCAAGGAGGUAAGCUUCUGCCCAGUGGUAAGCAAGUACUGUUGGCAGGAGCAGUCCUCCGAGGAGCCCACCGAGGACCAGUUCCACUC---CAGCGAGGGCG ..((((((((......))))..((((....(((((((((.((..(((......)))..))))))))))).((((((.(.((...)).).))))))........)))---).))))..... ( -51.20) >DroVir_CAF1 4192 120 + 1 GCUCGCGUAAUUCAUUCAGCAAGGAGGUGAGUUUCUGUCCAGUGGUGAGUAAGUAUUGCUGGCAGGAGCAAUCGUCGGAGGAGCCGCAAGAGGAGCACGAGCAGGCGAGCAGCGAUGAGG (((((((.....).....((.....(((..(((((((.((((..(((......)))..))))))))))).))).(((...(..((......))..).)))))..)))))).......... ( -40.60) >DroGri_CAF1 4801 114 + 1 GUUCGCGCAAUUCGUUCAGCAAGGAGGUGAGUUUCUGUCCGGUGGUGAGCAAGUAUUGCUGGCAGGAGCAGUCAUCGGAGGAGCCACAGGAGGAG------CAGGCGAGCAGCGACGAAG .(((((((..((((((..((.....((((((((((((.((((..(((......)))..)))))))))))..)))))((.....)).........)------).))))))..))).)))). ( -40.70) >DroWil_CAF1 2521 114 + 1 GAUCGAGAAAUUCCUUCAGCAAGGAGGUGAGUUUCUGUCCAGUCGUUAGCAAGUAUUGCUGGCAAGAGCAAUCCUCUGAAGAGCCACAAGAGGAGCAGGCAGA------CAGUGAUGAAG .((((((((((((((((......)))).)))))))).....((((((((((.....)))))))....((..((((((...........))))))....)).))------)..)))).... ( -36.30) >DroMoj_CAF1 2618 120 + 1 GCUCACGUAAUUCGUUUAGCAAGGAGGUGAGCUUCUGUCCCGUGGUGAGUAAGUAUUGCUGGCAGGAGCAGUCGUCGGAGGAGCCGCAGGAGGAGCAUGAGCAGACGAGCAGCGAUGAGG .(((((((..(((((((.((...((.(...(((((((.((.(..(((......)))..).)))))))))...).))....(..((......))..)....)))))))))..))).)))). ( -40.50) >DroAna_CAF1 21151 117 + 1 GUUCCCGAAACUCAUUUAGCAAGGAAGUAAGCUUCUGUCCAGUGGUAAGCAAGUACUGUUGGCAGGAGCAGUCCUCCGAAGAGCCCCAGGAGGAGCACGGAAAUUC---UAGCGAGGGAA .((((((.....).....((.((((.....(((((((.((((..(((......)))..)))))))))))..((((((...........)))))).........)))---).))..))))) ( -41.60) >consensus GAUCGCGAAAUUCAUUCAGCAAGGAGGUGAGCUUCUGUCCAGUGGUGAGCAAGUAUUGCUGGCAGGAGCAGUCCUCGGAGGAGCCACAGGAGGAGCACGAACAGGC___CAGCGAUGAAG .(((((((..(((..(((.(.....).)))(((((((.((((..(((......)))..)))))))))))...........)))((......))..................))).)))). (-24.70 = -23.57 + -1.13)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:45:32 2006