| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
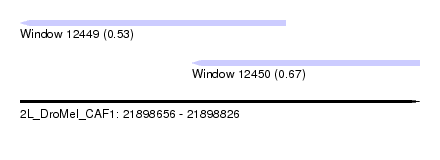
| Location | 21,898,656 – 21,898,826 |
| Length | 170 |
| Max. P | 0.665910 |

| Location | 21,898,656 – 21,898,769 |
|---|---|
| Length | 113 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 81.55 |
| Mean single sequence MFE | -24.46 |
| Consensus MFE | -16.95 |
| Energy contribution | -17.20 |
| Covariance contribution | 0.25 |
| Combinations/Pair | 1.20 |
| Mean z-score | -1.55 |
| Structure conservation index | 0.69 |
| SVM decision value | -0.01 |
| SVM RNA-class probability | 0.530799 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21898656 113 - 22407834 ACUGUACUAGUUCAUGGUUUCCAAAUAAGAGAAACCAGCACGGAAUUCCUAAACCAAA-------ACCUUUUCAGACCCUUGGCCCCGUAAGGCAUCACUUGCUACUCUUAGUAUCGGGC .(((.....((...((((((((......).)))))))..))((....)).........-------.......))).......((((.(((((......)))))(((.....)))..)))) ( -23.40) >DroSec_CAF1 188680 113 - 1 ACUGUACUAGUUCAUGGUUUCUACAUAAGAGAAACCAGAACGAAAUUCAUAAACCAAA-------CCCUUUCCAGACCCUUGUCCCCGUAAGGCAUCACUUGCUACUCUUAGUAUCUGGU .........((((.((((((((.......)))))))))))).................-------......(((((..((.......(((((......))))).......))..))))). ( -27.04) >DroSim_CAF1 132410 113 - 1 ACUGUACUAGUUCAUGGUUUCUACAUAAGAGAAACCAGAACGAAAUUCCUAAACCAAA-------CCCUUUCCAGACCCUUGUCCCCGUAAGGCAUCACUUGCUACUCUUAGUAUCGGGU .((((((((((((.((((((((.......))))))))))))((...............-------.........(((....)))...(((((......)))))...)).))))).))).. ( -26.30) >DroEre_CAF1 152877 118 - 1 ACUGAACUAGAUCACUGUUUCUAAAUUAGAGAAACGAGCACAGAAUUCCUAAGCCAAAAGCGAACACCUUGCCAGACCCCAAGCCAUGUUGGGCAUCACUUUCUACUCAUGGUAGA--AA .(((...........(((((((.......))))))).(((..(..(((((........)).)))..)..))))))..(((((......))))).......((((((.....)))))--). ( -21.10) >consensus ACUGUACUAGUUCAUGGUUUCUAAAUAAGAGAAACCAGAACGAAAUUCCUAAACCAAA_______ACCUUUCCAGACCCUUGGCCCCGUAAGGCAUCACUUGCUACUCUUAGUAUCGGGU ...((((((((((.((((((((.......)))))))))))).................................((.(((((......)))))..))............))))))..... (-16.95 = -17.20 + 0.25)



| Location | 21,898,729 – 21,898,826 |
|---|---|
| Length | 97 |
| Sequences | 3 |
| Columns | 119 |
| Reading direction | reverse |
| Mean pairwise identity | 77.97 |
| Mean single sequence MFE | -21.77 |
| Consensus MFE | -13.86 |
| Energy contribution | -14.87 |
| Covariance contribution | 1.01 |
| Combinations/Pair | 1.10 |
| Mean z-score | -1.68 |
| Structure conservation index | 0.64 |
| SVM decision value | 0.28 |
| SVM RNA-class probability | 0.665910 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21898729 97 - 22407834 --UUCU-----------C-GAAAUCGGUAUGAA--------UAGUUAAGAUUGAAUAGCUGCAAUUUAAUUUAAUGCAAACUGUACUAGUUCAUGGUUUCCAAAUAAGAGAAACCAGCA --....-----------.-(((..((((.((..--------..((((.((((((((.......)))))))))))).)).))))......))).((((((((......).)))))))... ( -17.00) >DroSec_CAF1 188753 118 - 1 AUCUCCCUUUUCGCUACGAGAAAUCGGUAUAAAA-AAAAUGUAGUUAAGAUUGAAUUGCUGCAAUUUCACUUAACAGCAACUGUACUAGUUCAUGGUUUCUACAUAAGAGAAACCAGAA ............(((((((....)))(((((...-....(((.((((((...((((((...))))))..)))))).)))..)))))))))((.((((((((.......)))))))))). ( -24.50) >DroSim_CAF1 132483 108 - 1 AUAUCU-----------GGUAAAUCGGUAUAAAAAAAAAUGUAGUUAAGAUUGAAUUGCUGCAAUUUCACUUAACAGCAACUGUACUAGUUCAUGGUUUCUACAUAAGAGAAACCAGAA ....((-----------((((...((((...........(((.((((((...((((((...))))))..)))))).)))))))))))))(((.((((((((.......))))))))))) ( -23.80) >consensus AUAUCU___________G_GAAAUCGGUAUAAAA_AAAAUGUAGUUAAGAUUGAAUUGCUGCAAUUUCACUUAACAGCAACUGUACUAGUUCAUGGUUUCUACAUAAGAGAAACCAGAA .....................(((..(((((........(((.((((((...((((((...))))))..)))))).)))..)))))..)))..((((((((.......))))))))... (-13.86 = -14.87 + 1.01)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:21 2006