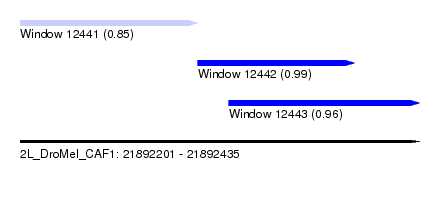
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 21,892,201 – 21,892,435 |
| Length | 234 |
| Max. P | 0.985579 |

| Location | 21,892,201 – 21,892,305 |
|---|---|
| Length | 104 |
| Sequences | 5 |
| Columns | 112 |
| Reading direction | forward |
| Mean pairwise identity | 93.36 |
| Mean single sequence MFE | -17.54 |
| Consensus MFE | -16.88 |
| Energy contribution | -16.88 |
| Covariance contribution | 0.00 |
| Combinations/Pair | 1.00 |
| Mean z-score | -1.22 |
| Structure conservation index | 0.96 |
| SVM decision value | 0.78 |
| SVM RNA-class probability | 0.849581 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
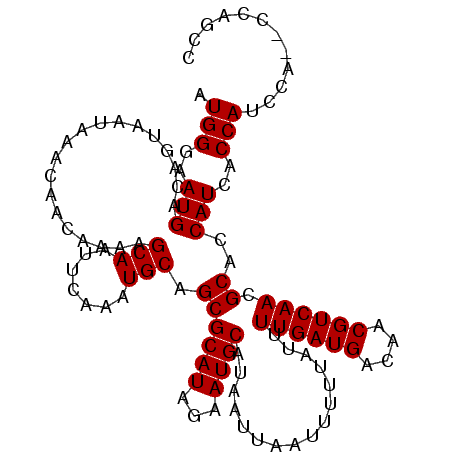
>2L_DroMel_CAF1 21892201 104 + 22407834 AUGGGAAUGACAGUAAUAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCAUCCA-------- ((((..(((...................(((.......))).((((((...))))................((((((....)))))).))..)))..))))...-------- ( -17.20) >DroSec_CAF1 181898 112 + 1 AUGGGAAUGACAGUAAUAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCAUCCAGCCCAGCC .((((...((..((..............(((.......))).((((((...))))................((((((....)))))).))......))..))...))))... ( -18.00) >DroSim_CAF1 125630 112 + 1 AUGGGAAUGACAGUAAUAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCAUCCAGCCCAGCC .((((...((..((..............(((.......))).((((((...))))................((((((....)))))).))......))..))...))))... ( -18.00) >DroYak_CAF1 139744 103 + 1 AUGGGAAUGACAGUAAUAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCAU---------CC ((((..(((...................(((.......))).((((((...))))................((((((....)))))).))..)))..))))---------.. ( -17.20) >DroAna_CAF1 126425 112 + 1 AUGGGAAUGACAGUAACAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCACCCCAACCAGUC .((((..(((..((..............(((.......))).((((((...))))................((((((....)))))).))))..)))....))))....... ( -17.30) >consensus AUGGGAAUGACAGUAAUAAACAACAAAAGCAAUUCAAAUGCAGCGCAUAGAAUGCAUAAUUAAUUUUUAUUUUGAUGACAACGUCAACGCACCAUCACCAUCCA__CCAGCC .(((..(((...................(((.......))).((((((...))))................((((((....)))))).))..)))..)))............ (-16.88 = -16.88 + 0.00)

| Location | 21,892,305 – 21,892,397 |
|---|---|
| Length | 92 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 67.82 |
| Mean single sequence MFE | -26.14 |
| Consensus MFE | -13.01 |
| Energy contribution | -13.33 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.27 |
| Structure conservation index | 0.50 |
| SVM decision value | 2.01 |
| SVM RNA-class probability | 0.985579 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
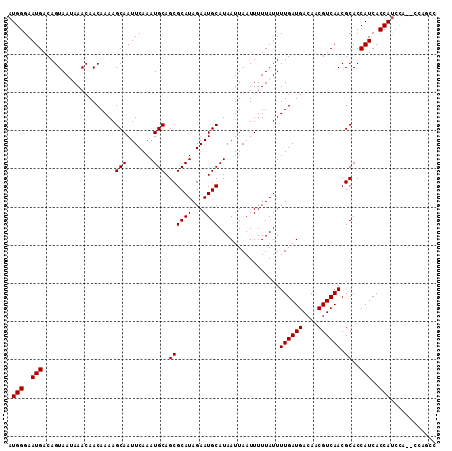
>2L_DroMel_CAF1 21892305 92 + 22407834 ----------------------GCCACCCAUUUUUCCAAA-ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCAGUGCAAUUAUCCUGUAUUGGACUUAUUUAC-GUACAUA----UGUAUG ----------------------(((((.............-.......)))))(((((....)))))((((((((((.......)))))))))).......(-((((...----.))))) ( -24.95) >DroEre_CAF1 145898 96 + 1 AGUCCAG------------CCCGUCGCCCUUUUUCACAAA-ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACUUAUGUAU-UUACAUA---------- (((((((------------.....................-.......((((((((((....)))))).))))((((.......)))))))))))(((((..-..)))))---------- ( -21.40) >DroYak_CAF1 139847 98 + 1 AGCCCAG------------CCAGCUGCCCAUUUUCCCAAA-ACAAUUCGUGCUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACAUAUGUAU-GUACAUA--------UG .(((.((------------(((((................-.........)))).)))....)))..(((((.((((.......)))).)))))((((((..-..)))))--------). ( -25.11) >DroAna_CAF1 126537 118 + 1 CCCGCGAUAAACUGCUCCACCA-CCUACCAUUUUUCCAAACACAUCCUCUGGGGUGCUCUUUGGCAUGUCCACUGCAAUUAUUUUGGAUUGGAUAU-UGUAUUGGACAAACGUAUGUGGA ...(((......)))....(((-(.(((.......(((((..(((((....)))))...)))))..((((((.((((((.((((......))))))-)))).))))))...))).)))). ( -33.10) >consensus AGCCCAG____________CCAGCCGCCCAUUUUCCCAAA_ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACAUAUGUAU_GUACAUA________UG ................................................(((..(((((....)))))(((((.((((.......)))).)))))............)))........... (-13.01 = -13.33 + 0.31)



| Location | 21,892,323 – 21,892,435 |
|---|---|
| Length | 112 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 68.97 |
| Mean single sequence MFE | -29.67 |
| Consensus MFE | -13.01 |
| Energy contribution | -13.33 |
| Covariance contribution | 0.31 |
| Combinations/Pair | 1.06 |
| Mean z-score | -2.49 |
| Structure conservation index | 0.44 |
| SVM decision value | 1.51 |
| SVM RNA-class probability | 0.959883 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
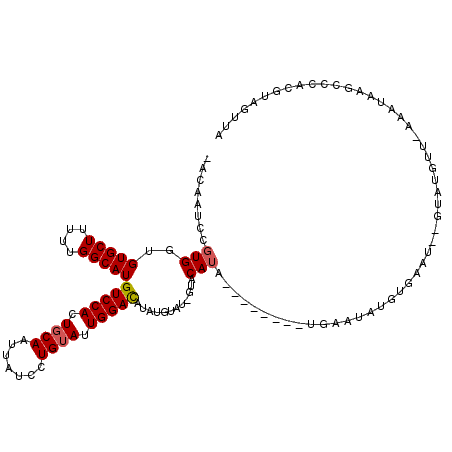
>2L_DroMel_CAF1 21892323 112 + 22407834 -ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCAGUGCAAUUAUCCUGUAUUGGACUUAUUUAC-GUACAUA----UGUAUGAAUAUCUGUAU--GCAUGUUAAAAUAAGCCAACGUAGUUA -......((((((.....(((((((((((((((((((.......)))))))))).......(-(((((.(----(((....)))).)))))--).)))))))))...))).)))...... ( -31.90) >DroEre_CAF1 145926 106 + 1 -ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACUUAUGUAU-UUACAUA------------UAUGUGAAUUGGAAUGCUCAAAUAAGCCCAGGUAGUUA -.....((.(((...(((((((((((((((((.((((.......)))).)))))......((-(((((..------------..)))))))....)))).)))).)))))))))...... ( -26.70) >DroYak_CAF1 139875 110 + 1 -ACAAUUCGUGCUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACAUAUGUAU-GUACAUA--------UGCAUAUGUGAAUUGGUAUGUGUAAAUACGCCCACGAAGUUA -....((((((...........((((((((((.((((.......)))).)))))))..((((-.((((((--------(((..((....))..))))))))).))))))))))))).... ( -34.00) >DroAna_CAF1 126576 101 + 1 CACAUCCUCUGGGGUGCUCUUUGGCAUGUCCACUGCAAUUAUUUUGGAUUGGAUAU-UGUAUUGGACAAACGUAUGUGGAAAUACGUAUAU--AUAC----------------AUAGUAA ..(((((....)))))..((.((...((((((.((((((.((((......))))))-)))).)))))).((((((......))))))....--...)----------------).))... ( -26.10) >consensus _ACAAUCCGUGGUGUGCUUUUUGGCAUGUCCACUGCAAUUAUCCUGUAUUGGACAUAUGUAU_GUACAUA________UGAAUAUGUGAAU__GUAUGUU_AAAUAAGCCCACGUAGUUA ........(((..(((((....)))))(((((.((((.......)))).)))))............)))................................................... (-13.01 = -13.33 + 0.31)

Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:44:14 2006