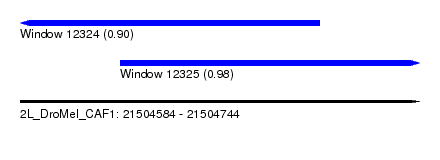

| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 21,504,584 – 21,504,744 |
| Length | 160 |
| Max. P | 0.982379 |

| Location | 21,504,584 – 21,504,704 |
|---|---|
| Length | 120 |
| Sequences | 4 |
| Columns | 120 |
| Reading direction | reverse |
| Mean pairwise identity | 85.63 |
| Mean single sequence MFE | -36.54 |
| Consensus MFE | -29.56 |
| Energy contribution | -28.88 |
| Covariance contribution | -0.69 |
| Combinations/Pair | 1.16 |
| Mean z-score | -2.03 |
| Structure conservation index | 0.81 |
| SVM decision value | 1.04 |
| SVM RNA-class probability | 0.904568 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21504584 120 - 22407834 UAAAUUUUUAAUGGAAGCGUCCACCAUUUGCUGAGUUGGCGGAUGUGACGGCGUCGCAGAGGCUUUCUUUGCCUUUGUGCCAGCAGAUCCAGAUGCCUUUUUUUGGACCACUUUCUUCUC ............(((((.((((((((((((((.....)))))))).)..((((((((((((((.......))))))))(....).......))))))......)))))..)))))..... ( -39.60) >DroSec_CAF1 149558 114 - 1 UAAAUUUUUAAUAGAAGCGUCCACCAUUUGCUGAGUUGGCGGAUGUGACGGCGUCGCAGUGGCUUUCUUAGCCUUUC------CGGAUCCAGAUGCCUUUUUUUGGACCACUUUCUUCUC ............(((((............((((((..(((.(.((((((...)))))).).)))..)))))).....------.((.((((((........)))))))).....))))). ( -32.20) >DroSim_CAF1 117108 120 - 1 UAAAUUUUUAAUAGAAGCGUCCACCAUUUGCUGAGUUGGCGGAUGUGACGGCGUCGCCGUGGCUUUCUUAGCCUUUGUGCCAGCGGAUCCAGAUGCCUUUUUUUGGACCACUUUCUUCUU ............(((((.((((((((((((((.....)))))))).)..((((((.(((((((...............)))).))).....))))))......)))))......))))). ( -35.86) >DroEre_CAF1 182528 104 - 1 -------------AAGGCGUCUACCAUUUGCUGAGUUGGCGGAUGUGACGGCGUCGCAGUGGCUUUCUUUGCCUUUGUG---GCAGAUGUAGAUGCCUUUUUUGGGGCCACUUUCUUCUC -------------(((((((((((((((((((.....)))))))).......(((((((.(((.......))).)))))---))....)))))))))))..................... ( -38.50) >consensus UAAAUUUUUAAUAGAAGCGUCCACCAUUUGCUGAGUUGGCGGAUGUGACGGCGUCGCAGUGGCUUUCUUAGCCUUUGUG___GCAGAUCCAGAUGCCUUUUUUUGGACCACUUUCUUCUC .............((((.((((.(((((((((.....)))))))).)..((((((.....(((.......)))(((((....)))))....)))))).......))))..))))...... (-29.56 = -28.88 + -0.69)

| Location | 21,504,624 – 21,504,744 |
|---|---|
| Length | 120 |
| Sequences | 3 |
| Columns | 120 |
| Reading direction | forward |
| Mean pairwise identity | 94.44 |
| Mean single sequence MFE | -34.33 |
| Consensus MFE | -32.29 |
| Energy contribution | -32.40 |
| Covariance contribution | 0.11 |
| Combinations/Pair | 1.03 |
| Mean z-score | -2.95 |
| Structure conservation index | 0.94 |
| SVM decision value | 1.91 |
| SVM RNA-class probability | 0.982379 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21504624 120 + 22407834 GCACAAAGGCAAAGAAAGCCUCUGCGACGCCGUCACAUCCGCCAACUCAGCAAAUGGUGGACGCUUCCAUUAAAAAUUUAAAGGAACGUGGCGGUUCAUCACUUCUGGCAAUCAAAAAAU ((.((.((((.......)))).)).((.((((((((.(((((((..........))))))).(.((((.((((....)))).)))))))))))))))..........))........... ( -35.80) >DroSec_CAF1 149595 117 + 1 ---GAAAGGCUAAGAAAGCCACUGCGACGCCGUCACAUCCGCCAACUCAGCAAAUGGUGGACGCUUCUAUUAAAAAUUUAAAGGAACGUGGCGGUUCAUCACUUCUUGCAAUAAAAAAAU ---.....((.(((((.((....))((.((((((((.(((((((..........))))))).(.((((.((((....)))).))))))))))))))).....)))))))........... ( -32.10) >DroSim_CAF1 117148 120 + 1 GCACAAAGGCUAAGAAAGCCACGGCGACGCCGUCACAUCCGCCAACUCAGCAAAUGGUGGACGCUUCUAUUAAAAAUUUAAAGGAACGUGGCGGUUCAUCACUUCUUGCAAUAAAAAAAU (((....((((.....))))..(..((.((((((((.(((((((..........))))))).(.((((.((((....)))).)))))))))))))))..)......)))........... ( -35.10) >consensus GCACAAAGGCUAAGAAAGCCACUGCGACGCCGUCACAUCCGCCAACUCAGCAAAUGGUGGACGCUUCUAUUAAAAAUUUAAAGGAACGUGGCGGUUCAUCACUUCUUGCAAUAAAAAAAU ........((.(((((.((....))((.((((((((.(((((((..........))))))).(.((((.((((....)))).))))))))))))))).....)))))))........... (-32.29 = -32.40 + 0.11)



Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:42:26 2006