
| Sequence ID | 2L_DroMel_CAF1 |
|---|---|
| Location | 21,327,566 – 21,327,682 |
| Length | 116 |
| Max. P | 0.946426 |

| Location | 21,327,566 – 21,327,662 |
|---|---|
| Length | 96 |
| Sequences | 6 |
| Columns | 99 |
| Reading direction | reverse |
| Mean pairwise identity | 73.13 |
| Mean single sequence MFE | -45.28 |
| Consensus MFE | -16.40 |
| Energy contribution | -18.10 |
| Covariance contribution | 1.70 |
| Combinations/Pair | 1.50 |
| Mean z-score | -2.43 |
| Structure conservation index | 0.36 |
| SVM decision value | 1.35 |
| SVM RNA-class probability | 0.945202 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
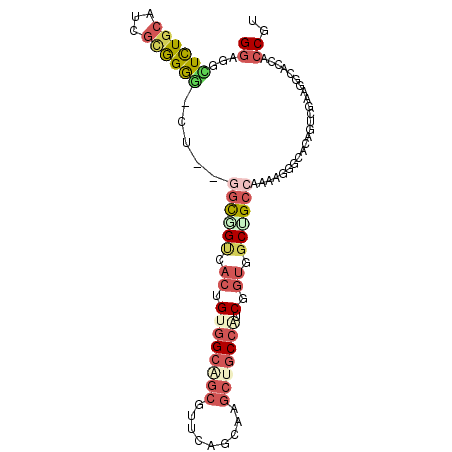
>2L_DroMel_CAF1 21327566 96 - 22407834 GGAUGCUCUGCAUCGCGGGG-CU--GGUUGUCACUGUGGCAGCGUUCAGUAGGCUGCCGUCGGUGGCUGCCAAAAUGGCGCAGUCAAAGGCACCUCCGC (((.(((((((...))))))-)(--(((.((((((((((((((.(.....).))))))).))))))).))))....((.((........)).))))).. ( -45.70) >DroVir_CAF1 31766 96 - 1 GCAGUGUUUGCAUUGUGGGC-GU--GGCAGCAACAGUUGCCGCACUCAAUAAGCUGCCAUCGCUGGCAGCGAGCAGCGCACAGUCAUAGGCACCACCGC ...(((((((.((((((.((-((--((((((....)))))))).........(((((((....))))))).....)).))))))..)))))))...... ( -43.70) >DroEre_CAF1 26343 96 - 1 GGAUGCUCUGCAUCGCGGGG-CU--GGCGGUCACUGUGGCAGCGUUCAGCAAGCUGCCAUCGGUGGCUGCCAAAAGGGCACAGUCGAAGGCGCCUCCGU .(((((...)))))((((((-((--((((((((((((((((((.........))))))).))))))))))))..........((.....))))).)))) ( -51.70) >DroYak_CAF1 30098 96 - 1 GGAUGCUCUGCAUCGCGGGG-CU--CGCGGUGACUGUGGCAGCGUUCAGCAAGCUGCCAUCAGUGGCUGCCAAAAGGGCACAGUCGAAUGCGCCACCGU .(((((...)))))((((((-(.--.(((((.(((((((((((.........))))))).)))).))))).......(((........)))))).)))) ( -45.70) >DroAna_CAF1 20630 99 - 1 GGAGGUUCUGCAUCGAGGAGGCUGUGCUUGCCACAGCGGAGGCGUUCAGCAGGCUGCCAUCGGUGGCGGCCAGGAGCGCACAGUCGUAGGAUGCCCCGU ((.((((((((.((...))((((((((..(((........)))((((....((((((((....))))))))..)))))))))))))))))..))))).. ( -46.90) >DroPer_CAF1 22847 96 - 1 GGAGACUCUGCAUCACAGGA-UU--GGCCGAAACGGCGGCUGCAUUCAGCAGAGUGCCAUCGGCGGCGGCCAACAGAGCGCAAUCGUAGGCCCCACCAU ((..((((((((((....))-).--(((((......))))).......))))))).))...((.((.((((.((.((......)))).)))))).)).. ( -38.00) >consensus GGAGGCUCUGCAUCGCGGGG_CU__GGCGGUCACUGUGGCAGCGUUCAGCAAGCUGCCAUCGGUGGCUGCCAAAAGGGCACAGUCGAAGGCACCACCGU ((...((((((...)))))).....((((((.((.((((((((.........))))))).).)).))))))........................)).. (-16.40 = -18.10 + 1.70)


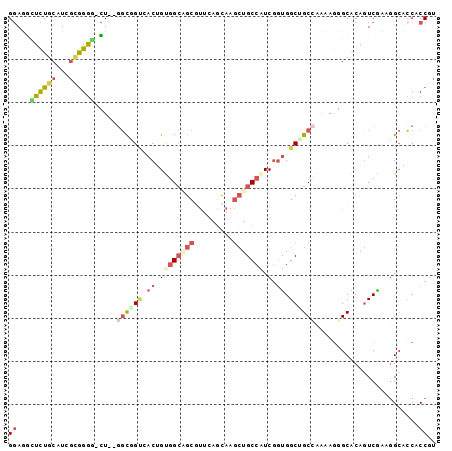
| Location | 21,327,585 – 21,327,682 |
|---|---|
| Length | 97 |
| Sequences | 6 |
| Columns | 100 |
| Reading direction | reverse |
| Mean pairwise identity | 75.78 |
| Mean single sequence MFE | -43.35 |
| Consensus MFE | -21.71 |
| Energy contribution | -22.27 |
| Covariance contribution | 0.56 |
| Combinations/Pair | 1.41 |
| Mean z-score | -2.29 |
| Structure conservation index | 0.50 |
| SVM decision value | 1.36 |
| SVM RNA-class probability | 0.946426 |
| Prediction | RNA |
Download alignment: ClustalW | MAF
>2L_DroMel_CAF1 21327585 97 - 22407834 CAAGUGUAUGGCUUGUGAUUGGAUGCUCUGCAUCGCGGGG-CU--GGUUGUCACUGUGGCAGCGUUCAGUAGGCUGCCGUCGGUGGCUGCCAAAAUGGCG .((((.....))))((.(((....(((((((...))))))-)(--(((.((((((((((((((.(.....).))))))).))))))).)))).))).)). ( -42.20) >DroPse_CAF1 21026 97 - 1 CAUAUGCAUCGCCUGGGGCUGGAGACUCUGCAUCACAGGA-UU--GGCCGAAACGGCGGCUGCAUUCAGCAGAGUGCCAUCGGCGGCGGCCAACAGAGCG .....((.((.((((((((.(....)...)).)).)))).-((--(((((...((.(((..((((((....))))))..))).)).)))))))..)))). ( -40.70) >DroEre_CAF1 26362 97 - 1 CAAGUGUAUGGCUUGUGAUUGGAUGCUCUGCAUCGCGGGG-CU--GGCGGUCACUGUGGCAGCGUUCAGCAAGCUGCCAUCGGUGGCUGCCAAAAGGGCA ..........((((..........(((((((...))))))-)(--((((((((((((((((((.........))))))).))))))))))))...)))). ( -48.10) >DroYak_CAF1 30117 97 - 1 CAAGUGUAUGGCUUGUGAUUGGAUGCUCUGCAUCGCGGGG-CU--CGCGGUGACUGUGGCAGCGUUCAGCAAGCUGCCAUCAGUGGCUGCCAAAAGGGCA ..........((((.((....((.(((((((...))))))-))--)(((((.(((((((((((.........))))))).)))).)))))))...)))). ( -43.20) >DroAna_CAF1 20649 100 - 1 UAUGUGCAUGCUCUGUGGCUGGAGGUUCUGCAUCGAGGAGGCUGUGCUUGCCACAGCGGAGGCGUUCAGCAGGCUGCCAUCGGUGGCGGCCAGGAGCGCA ....(((..(((((...((((((.(((((((.((...))(((.......)))...))))).)).)))))).((((((((....)))))))).)))))))) ( -45.20) >DroPer_CAF1 22866 97 - 1 CAUAUGCAUCGCCUGGGGCUGGAGACUCUGCAUCACAGGA-UU--GGCCGAAACGGCGGCUGCAUUCAGCAGAGUGCCAUCGGCGGCGGCCAACAGAGCG .....((.((.((((((((.(....)...)).)).)))).-((--(((((...((.(((..((((((....))))))..))).)).)))))))..)))). ( -40.70) >consensus CAAGUGCAUGGCCUGUGACUGGAGGCUCUGCAUCGCGGGG_CU__GGCCGUCACUGCGGCAGCGUUCAGCAGGCUGCCAUCGGUGGCGGCCAAAAGGGCA ....(((....(((((((.((.(.....).)))))))))......((((((.((...((((((.........))))))....)).))))))......))) (-21.71 = -22.27 + 0.56)

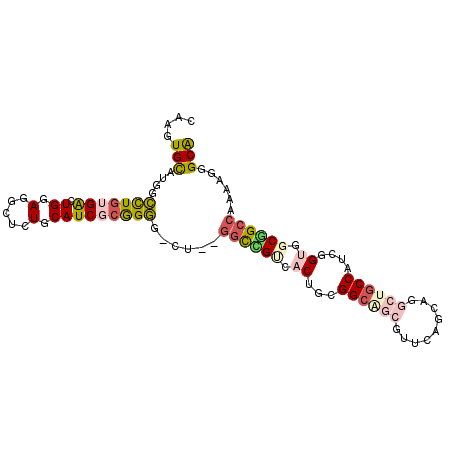
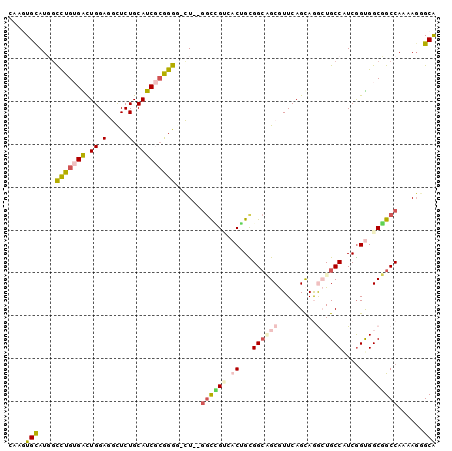
Generated by rnazCluster.pl (part of RNAz 1.0) on Mon Dec 4 12:42:07 2006